Cambridge University 2006
From 2006.igem.org
| - Autonomous Pattern Formation between Bacterial Populations by Reciprocal Communication - |
-
Project Summary
Our entry for the iGEM 2006 competition sought to genetically engineer two types of bacteria such that they would autonomously form patterns in swimming agar. We studied the dynamics of pattern formation between bacterial populations. In particular, we investigated the interaction of genetically engineered E. coli sender cells, producing an acyl homoserine lactone signal, and receiver cells in swimming agar. We then attempted to render these cells capable of reciprocal communication and sought to construct biological parts that would render possible the construction of a population based bi-stable switch. Furthermore, we modelled the behaviour of single cells as well as the interactive behaviour of populations of cells containing these genetic circuits. Our experiments verified that differential cell motility in combination with position-dependent gene expression has the potential to generate complex patterns.
Introduction
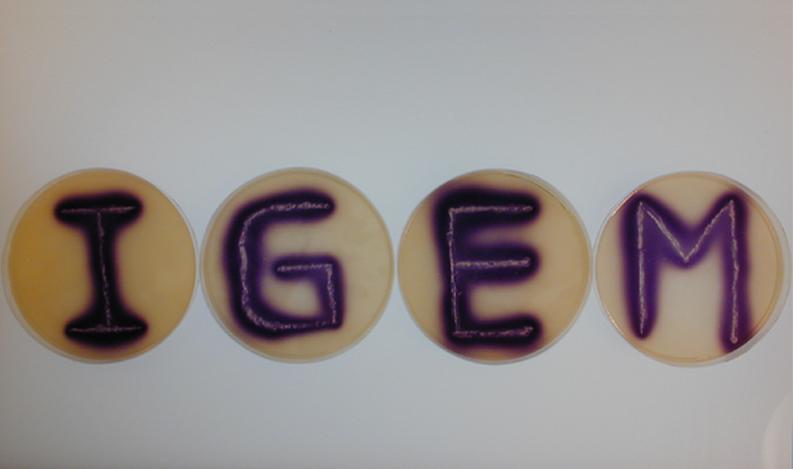
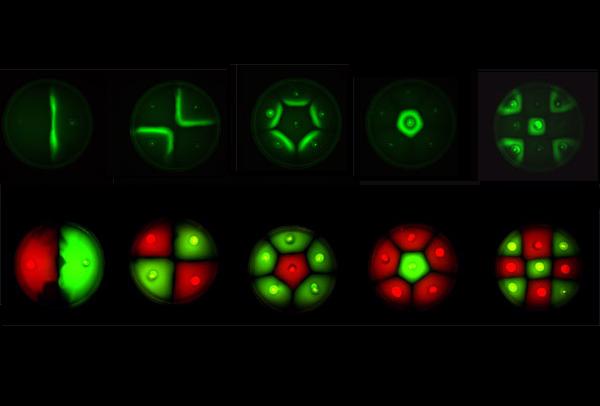
Position dependent gene expression is a critical aspect of the behaviour of multi-cellular organisms and requires a complex series of interactions to occur between cell types. We used Escherichia coli as the organism of choice for our experiments and observed how differential cell motility can in itself lead to complex pattern formation (Figure 1.1). We genetically engineered an E. coli cell population to render it capable of producing an acyl-homoserine lactone signal (sender cells). We verified that the cells were producing lactones using the CVO26 plate assay described by McClean et al. (Figure 1.2). A second E. coli population was rendered capable of responding to this acyl homoserine lactone signal (receiver cells). Adapting the experiments of Weiss et al., using cell motility, rather than a differential response to AHL concentrations as a way to define zones of response we noted how the interaction of sender and receiver cell populations on a swimming plate leads to complex pattern formation (Figure 1.3). For both experiments mentioned above we used parts from the MIT Registry of Standard Biological Parts, thus characterizing their function in this novel fashion. Thereafter, we proceeded to make the genetic constructs described in “Biological Parts” below. Equipping highly motile strains such as E. coli MC1000 with AHL-mediated autoinducing systems based on Vibrio fischeri luxI/luxR and Pseudomonas aeruginosa lasI/lasR cassettes would allow the amplification of a response to an AHL signal and its propagation. Exploiting the uncoupled nature of sender and receiver entities, both AHL systems can be combined to give two-way communication pathways. The recursive behaviour of cells being both sender and receiver at the same time has the potential to create patterns that are robust and more complex in nature. Although we were unable to construct the complete system, we were able to construct auto-inducing luxI/R and lasI/R cassettes. For some of our constructions we used the method of 3-Antibiotic assembly, described by Tom Knight et al. We were able to purify standard plasmid backbone vectors for this assembly method and also to get it to work for some of the constructions as evidenced by the parts submitted. We also managed to model the behaviour of cells within populations containing these systems. Further, we believe that in the future it may be possible to equip E. coli populations with more complex genetic circuits that would allow two interspersed subpopulations of isogenic bacteria to battle for dominance on a swimming plate. This is described in detail in "Future Directions".
|
| ||||||||