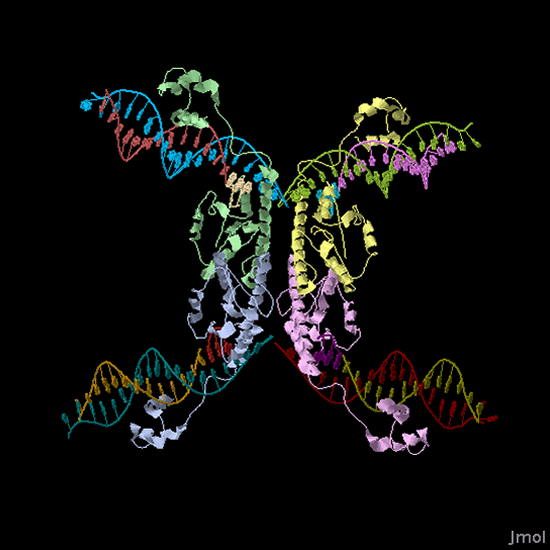Davidson 2006
From 2006.igem.org
| Line 39: | Line 39: | ||
<big>'''Methods and Results'''</big><br> | <big>'''Methods and Results'''</big><br> | ||
| + | The following is an outline of what the Davidson team will present at iGEM 2006<br><br> | ||
'''Basic parts''': Parts used in this project were designed by the [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Davidson Davidson] and [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Missouri Missouri Western] iGEM teams<br> | '''Basic parts''': Parts used in this project were designed by the [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Davidson Davidson] and [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Missouri Missouri Western] iGEM teams<br> | ||
'''Modeling''' | '''Modeling''' | ||
| Line 44: | Line 45: | ||
*Using modeling to choose which families of unsolved pancake stacks to start with | *Using modeling to choose which families of unsolved pancake stacks to start with | ||
'''Building the Biological System''' | '''Building the Biological System''' | ||
| - | *Single | + | *"Single pancake" constructs and initial observations |
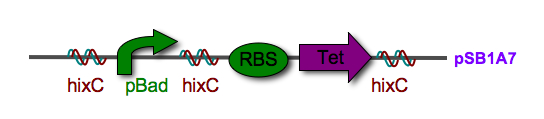
| - | * | + | *Read-through from the carrier vector backbone leads to uncontrolled Tet expression and uncontrolled flipping |
| - | *New [http://partsregistry.org/Part:BBa_J31009 pSB1A7] vector | + | *New [http://partsregistry.org/Part:BBa_J31009 pSB1A7] vector insulates parts from read-through, but is not compatible with parts carrying double terminators |
| - | * | + | *Solution: designing pancake constructs without double terminators |
| - | *Two pancake constructs | + | *"Two-pancake" constructs |
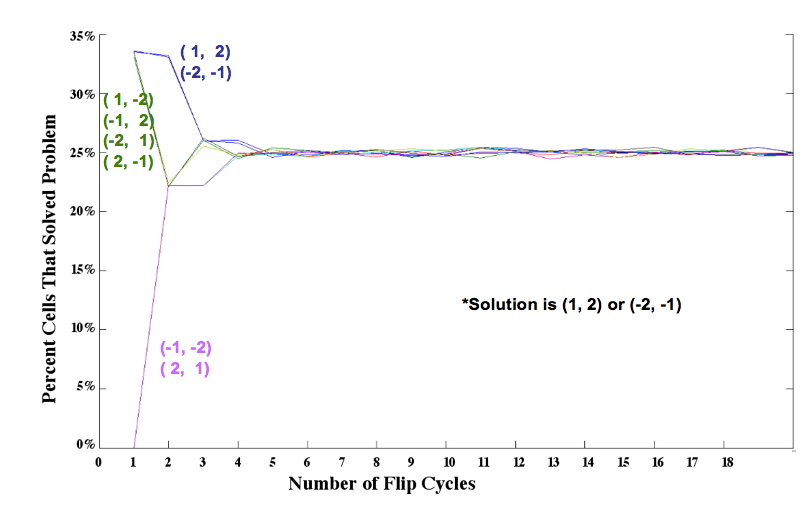
| - | * | + | *Dealing with biological equivalence: distinguishing 1,2 from -2,-1 using an RFP reporter |
| + | *Updated design | ||
<big>'''Conclusions'''</big><br> | <big>'''Conclusions'''</big><br> | ||
| - | *'''Consequences of devices''': | + | *'''Consequences of devices''' |
| + | ** Practical: proof-of-concept for bacterial computers, data storage | ||
| + | ** Basic research: transgene rearrangement in vivo, insights into evolution that has occurred via DNA rearrangements | ||
*'''Next steps''': can solve problem but need control over kinetics | *'''Next steps''': can solve problem but need control over kinetics | ||
*'''Lessons learned''': | *'''Lessons learned''': | ||
Revision as of 00:23, 29 October 2006
 Project Overview Davidson Parts Team Members Tools and Resources Check out our Official Team Photo |
Our goal is to genetically engineer a biological system that can compute solutions to a puzzle called the burnt pancake problem. The EHOP computer is a proof of concept for computing in vivo, with implications for future data storage devices and transgenic systems. Our work was done in collaboration with the Missouri Western iGEM Team and an undergraduate research fellow from Hampton University.
The Burnt Pancake Problem
The pancake problem is a puzzle in which a scrambled series of units (or stack of pancakes) must be shuffled into the correct order and orientation. You can try solving a simple version of the pancake problem yourself.
| In the burnt pancake problem, each pancake is given an orientation by burning one side. Figure 1 shows a scrambled stack of burnt pancakes. To solve the problem, every unit, or pancake, must be placed in the proper order (largest on bottom, smallest on top) and in the proper orientation (burnt side down, golden side up). The pancakes are flipped with two spatulas: one to lift pancakes off the top of the stack, the other to flip part (or all) of the remaining stack of pancakes. The pancakes lifted by the first spatula are returned to the top of the stack without being flipped. You can watch Media:burnt_pancake.ogg a movie of the stack in Figure 1 being sorted to see how the puzzle is solved.
Building the Biological System
TEAM MEMBERS
Faculty
TOOLS AND RESOURCES
White Board Biology Tools (Wet Bench)
Math Tools
Bio-Math Tools
Assembly Plans Progress |