Mathematical Modeling
From 2006.igem.org
Contents
|
Assumptions
- All zincfingers have the same affinity (K) and the same cooperativity (n).
- The input signal S consists of two parts, one of them activates the promoter to two of the zincfingers, the other represses the promotor to the other two zincfingers.
- The input signal S is assumed to be a sine wave with breaks (times where it stays zero) in between.
- There is no stochastic noise included in the model.
- In the start state, zincfinger 1 is expressed.
Deriving a model
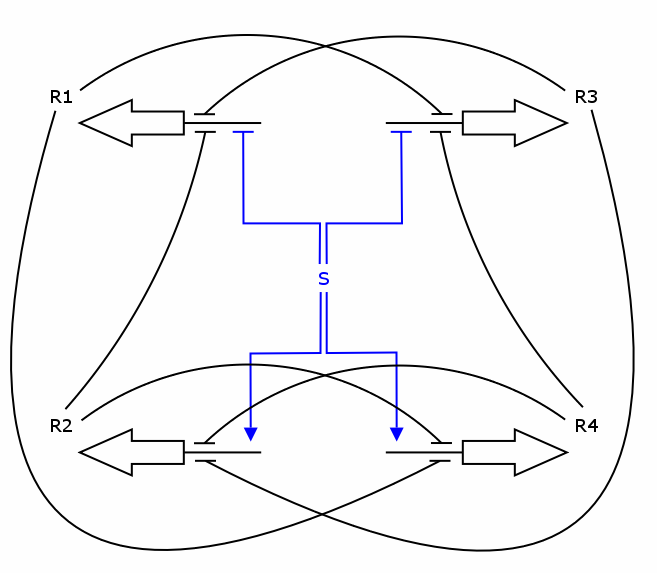
Recall the counter architecture:
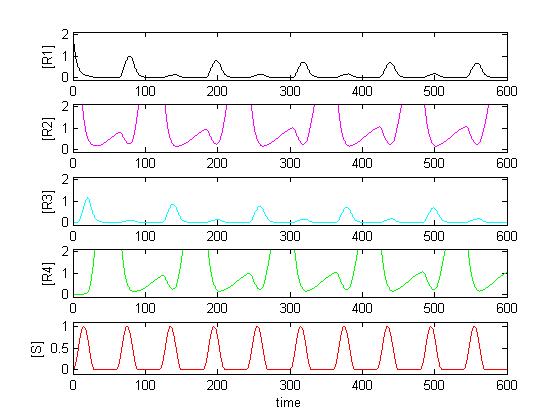
We want to describe the concentrations of zincfingers R1 to R4 over time given the input signal S. We will set up a deterministic model using ordinary differential equations (ODEs), as this approach is commonly used in modeling gene networks.
General approach
We have to formulate for each gene product an ODE describing its rate of change. This rate is comprised of a term characterizing the synthesis rate and one characterizing the degradation rate.
We assume that the degradation rate of a protein depends linearly on its concentration, i.e.
deg(p) = -k_deg*p
where k_deg is a degradation constant.
As for the synthesis rate we have to consider that gene expression is controlled by regulatory proteins, either
in an activating or a repressing manner.
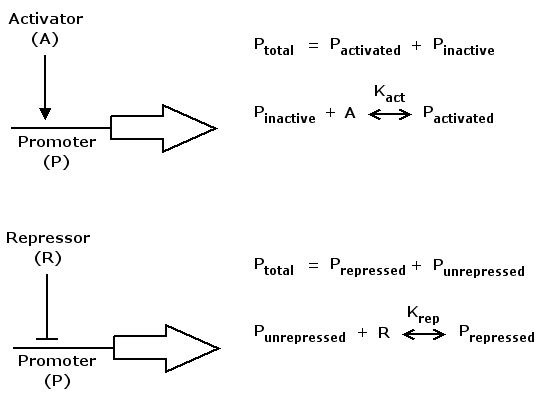
Mathematically, activation can be formulated in the following way
act(A) = (A/K_act)^n / (1+(A/K_act)^n)
where A is a regulatory protein activating gene expression, K_act is a constant characterizing promoter-activator affinity (see figure above) and n describes the cooperativity of the activator. act(A) is in the range of [0 .. 1] and increases monotonically with increasing A.
Repression can be formulated accordingly
rep(R) = 1 / (1+(R/K_rep)^n)
where R is a regulatory protein repressing gene expression, K_rep and n are the equivalents of K_act and n in the activation case. rep(R) is in the range of [0 .. 1] and decreases monotonically with increasing R.
For the input signal S it needs to be taken into account that the two promoters (Pr and Prm) have different activity levels. Activation of S is therefore formulated as
act(S) = ((S/K_act)^n / (1+(S/K_act)^n) + P1/(P2-P1)) * (P2-P1)
repression as
rep(S) = (1 / (1+(S/K_rep)^n) + P3/(P4-P3)) * (P4-P3)
where act(S) is in the range of [P1 .. P2] and rep(S) in the range of [P3 .. P4]. act(S) and rep(S) therefore need to be scaled in such a way that they fall within the range of [0 .. 1]
To get a synthesis rate we can multiply both act() and rep() with a kinetic constant k that describes the maximum synthesis rate. So on one hand act() and rep() denote the synthesis rate relative to the maximum rate. On the other hand we can interpret them as probabilities. That is, the probability of any RNA polymerase molecule binding to the respective promoter and starting the protein synthesis process.
The total rate of change is then obtained by adding together the degradation and synthesis rates.
Application to the counter circuit
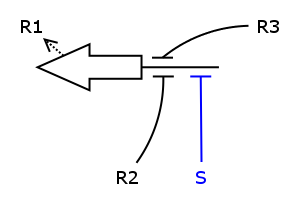
We now have all the building blocks we need to construct the ODEs for R1 to R4. Let's have a closer look at one of those Rs, say R1.
As one can see, the production of R1 depends on S, R2 and R3, i.e. S activates the production and R2/R3 repress it. We assume the individual repressors and activators affect the synthesis of R1 independently from each other, which means we can use the expressions for activation and repression that we have already formulated for each regulator separately. As we stated above the values of act() and rep() can be interpreted as probabilities. So what we need to model is the probability of three independent events coinciding. As we know from probability theory we obtain the desired probability by multiplying the individual event probabilities. Gluing everything together we get
dR1/dt = k_syn_R1 * rep(S) * rep(R2) * rep(R3) - k_deg_R1 * R1
\_________________ _________________/ \_______ _______/
V V
synthesis rate degradation rate
Similarly we construct the other equations
dR2/dt = k_syn_R2 * act(S) * rep(R3) * rep(R4) - k_deg_R2 * R2 dR3/dt = k_syn_R3 * rep(S) * rep(R1) * rep(R4) - k_deg_R3 * R3 dR4/dt = k_syn_R4 * act(S) * rep(R1) * rep(R2) - k_deg_R4 * R4
Possible model improvements
Firstly, since the biological implementation of the circuit is split up into the Event_Processing_Device and 4 overlapping NOR modules of the 4-State_Device, a more detailed/sophisticated model would probably give a better approximation to the actual circuit.
Secondly, through measuring the Event_Processing_Device, a better approximation of the periodical shape of the input signal S could be found.
Sensitivity Analysis
Intro
The goal of sensitivity analysis is to find out to which parameters our model is sensitive. When some parameters are changed slightly, the system can respond anywhere between weakly and strongly. If it reacts strongly, i.e. the output of the system changes drastically, then the system is very sensitive to that parameter. If the system only changes its output slightly (or not at all) when a certain parameter is changed, the system is said to be insensitive to that parameter.
It goes without saying that especially the sensitive parameters need to be taken care of, as different settings of the insensitive parameters don't affect the output of the system much. Based on the sensitive parameters it can be decided which parameters should be tested.
Multiparametric Sensitivity Analysis (MPSA)
One method to find the sensitive parameters is the Multiparametric Sensitivity Analysis which is based on the Monte Carlo Method. The method is used as it is described in cho04a-sensitivity. It works as follows:
- The nominal parameters for the observed output f_obs are chosen. In our case, these parameters should be chosen such that the oscillation works correctly. The observed output serves as a reference for all the outputs that generated with varied input parameters.
- The observed output is calculated, which means that the differential equations need to be solved. In our case, the observed output is the concentration of zincfinger 4 (because ZF4 oscillates best for that set of nominal parameters).
- The parameters whose sensitivity is to be compared are chosen. In our case, only two parameters at a time are compared (because the computational effort increases exponentially with the number of parameters that are to be compared).
- For both chosen parameters, generate a series of independet random numbers that are uniformly distributed within the range of the respective parameter.
- For each set of random parameter values (i.e. one random value of parameter 1 and one random value from parameter 2, for every possible combination of the random values) solve the differential equations to get the perturbed output f_per.
- For each perturbed output calculate the corresponding objective function as follows: <math>f_{obj}(k) = \sum_{i=1}^q (f_{obs}(i) - f_{per}(i,k))^2</math> (why the hell doesn't this *f--* math thing work ?!?), which is the sum of square errors between the initial observed and the perturbed error.
- Set the acceptance criterion as the average of the objective function.
- A set of parameters is acceptable if it is greater than the criterion, else it is unacceptable.
- For both parameters, calculate the frequency distribution, i.e. count for every random number (from step 3) the number of objective functions that are acceptable or unacceptable, respectively (for example: for each random value of parameter 1 there is one objective function for every random value of parameter 2).
- For both parameters, calculate the accumulated frequency distributions for the acceptable und the unacceptable cases (i.e. add them together incrementally).
- To measure the sensitivity of the parameters, calculate the correlation coefficient between the cumulative distributions for the acceptable and the unacceptable cases.
- Compare the correlation coefficients of the two parameters. The parameter with the lower correlation coefficient is more sensitive than the other.
Example
In this example the sensitivities of K (affinity of the zincfingers) and KS (affinity of the input signal S) are compared.
The nominal values and ranges are the same as in the chapter 'Results'.
Firstly, 20 random numbers for both K and KS are computed such that the numbers are in the uniformly distributed in the ranges of K and KS, respectively.
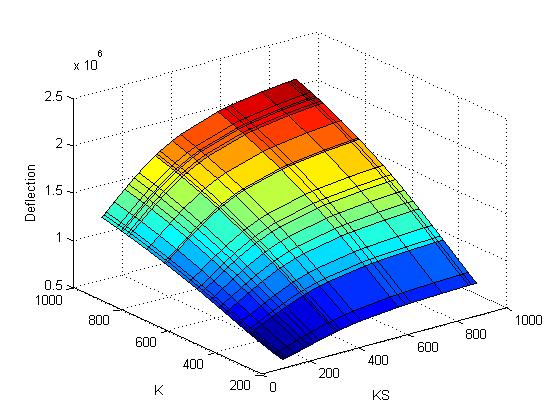
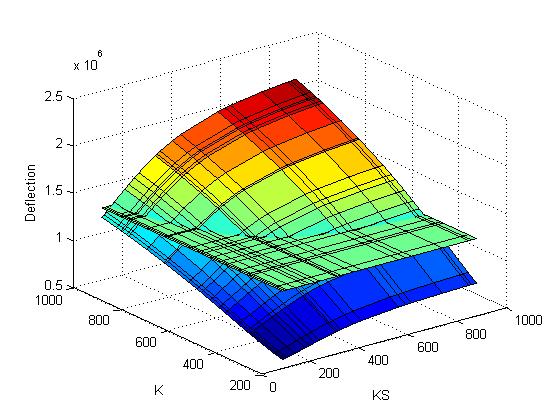
The objective function (Deflection) of each possible pair of values for K and KS are computed and plotted:
The criterion gets computed and added to the plot:
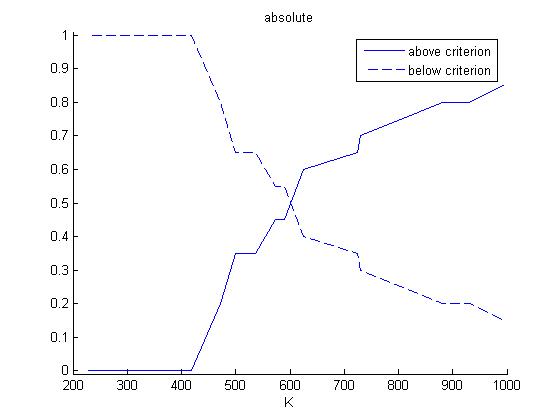
Calculation of the frequency distribution: For every one of the 20 values of K, there are 20 objective function values (one for each value of KS). For every value of K, the number of objective function values that are above and below the criterion, respectively, are counted. Then the numbers get normalized to 1 and plotted:
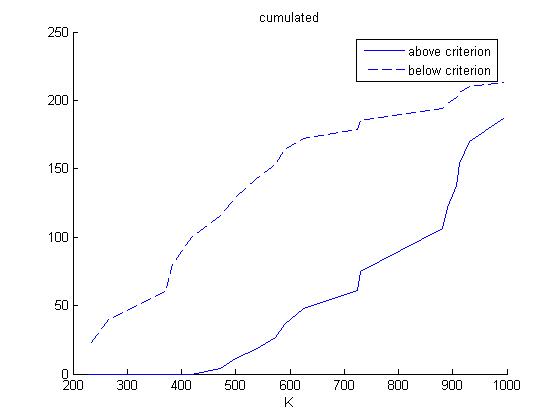
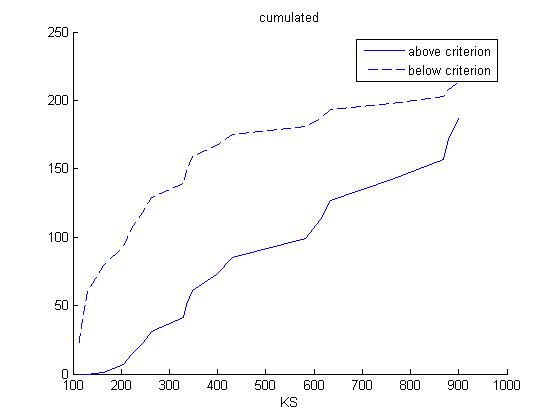
The non-normalized numbers now get accumulated:
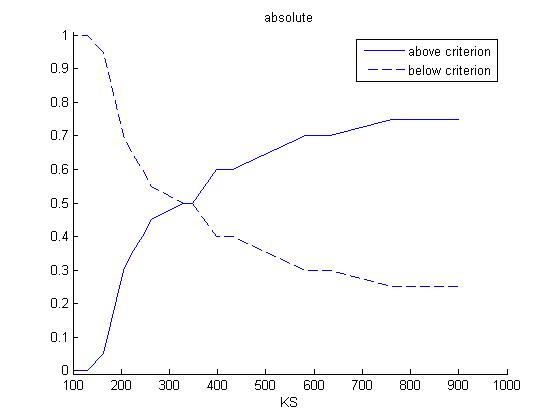
Then the same procedure is applied to every value of KS to calculate the frequency distribution of KS:
Finally, the correlation coefficient for both K and KS are computed. This correlation coefficient is the correlation between the cumulated frequency distribution of the plots 'above criterion' and 'below criterion' for each of the two parameters.
The correlation coefficient are 0.8325 for K and 0.9086 for KS. Hence, K is more sensitive than KS.
Results
6 Parameters were compared in pairs against each other. These Parameters were:
| Parameter: | Description: | Range: | Nominal value: |
| n | Cooperativity of the zincfingers | 2-4.5 | 3 |
| nS | Cooperativity of the input signal S | 2-4.5 | 3 |
| K | Affinity of the zincfingers | 66-150 | 100 |
| KS | Affinity of the input signal S | 66-150 | 100 |
| k | maximum synthesis rate of the zincfingers | 533-1200 | 800 |
| kd | degradation rate of the zincfingers | 0.66 - 1.5 | 1 |
| SA | maximum concentration of the input signal S | 333-750 | 500 |
The nominal values of the parameters were chosen in such a way that the counter oscillates in the desired manner. By applying MPSA the relative sensitivities of each possible pair of parameters were computed (see table below).
The ranges are chosen such that they go from (nominal value)/1.5 to (nominal value)*1.5. That way, it is ensured that the counter oscillates correctly for every varying pair of parameters.
The concentration of zincfinger 4 was chosen to be the output of the system.
For example n was compared against nS (first entry in table below). The MPSA method returns a correlation coefficient for each one of the two parameters. In our case the correlation coefficient of n was smaller than that of nS, which means that n is more sensitive than nS. To compute the relative sensitivity the correlation coefficient of nS was divided by the correlation coefficient of n, the result of which is given in the first entry in the table below.
In other words: Numerical value of table = (Correlation coefficient of Parameter 2)/(Correlation coefficient of Parameter 1).
Again in other words: If an entry in the table below is bigger than one, it means that Parameter1 is more sensitive than Parameter2. For example in the second least row (of kd) every entry is bigger than one which means that kd is more sensitive than any other parameter.
The Parameters 1 and 2 are those in the first column and row, respectively.
| Parameter2 | n | nS | K | KS | k | kd | SA | |
| Parameter1 | - | - | - | - | - | - | - | - |
| n | - | - | 0.617759261 | 0.744339192 | 0.67 | 0.6913 | 0.636148929 | 0.607330169 |
| nS | - | 1.618931425 | - | 0.924307575 | 0.527508226 | 0.59565 | 0.548246533 | 0.685923595 |
| K | - | 1.34360469 | 1.083029731 | - | 0.561914972 | 0.5997 | 0.709109014 | 0.787966708 |
| KS | - | 1.515065483 | 1.918471156 | 1.805039619 | - | 0.57877227 | 0.765449083 | 1.100975806 |
| k | - | 1.450549543 | 1.695672782 | 1.691273469 | 1.729421028 | - | 1.016717207 | 1.679752051 |
| kd | - | 1.581022928 | 1.824024904 | 1.414348333 | 1.314024447 | 0.983557673 | - | 1.669685461 |
| SA | - | 1.648169059 | 1.506232516 | 1.269584428 | 0.909901984 | 0.595410883 | 0.599110258 | - |
Note: The values for nS-K were contradictory. So for that pair, another pair of nominal values and ranges was chosen: 4 for nS (2.6 - 6) and 200 for K (133 - 300).
From left (most sensitive) to right (least sensitive), the parameters are as follows:
k kd KS SA K nS n
It can be seen that the maximum production rate and the degradation rate of the zincfingers are the most sensitive parameter (this doesn't mean that these parameters have to be big, it just means that a slight change in the degradation or maximum production rate has a big influence on the output).
It can also be seen that the input signal is more sensitive than the Zinc Fingers, and that the affinities are more sensitive than the cooperativities.
Results from cloning
- The activity of Pr and Prm were compared where no IPTG and therefore no cI was present. It seems that in this state Pr is about 3-6 times stronger than Prm. The other state (where IPTG and cI are present) has not been tested yet.
Which two Zincfingers should have the same affinity?
At the meeting yesterday, the question came up whether some zincfingers need to have the same affinity. Because as it is, we don't have four zincfingers with the same affinity. So I did some quick modeling, assuming that two ZFs have an affinity of 3 nmol and the other two an affinity of 30 nmol. As it turns out, the only constellation that works is if ZF1 and ZF3 have an affinity of 30 and ZF2 and ZF4 an affinity of 3:
Matlab Code
There are two packages that can be downloaded. The first package is the one that calculates the concentration of the zincfingers for the different parameters, the second one is for the sensitivity analysis.
For those who want to try out the Matlab code, please use the zipped version and not the *.m files on the wiki (because those have been adapted to fit into the wiki and hence contain mistakes).
[http://www.tik.ee.ethz.ch/%7ebarkows/counter.zip Zincfinger concentrations gezippt]
[http://people.ee.ethz.ch/~ulrichta/sensitivity.zip Sensitivity gezippt]
Simulation for people who don't know how to handle Matlab
- If you don't have Matlab on your computer, install it (unfortunately, it's payware).
- Save all the *.m files (Matlab-files) from the zipped Matlab Code to your computer.
- Open Matlab.
- Set the 'Current Directory' path (above the Matlab terminal) to the folder where you stored the *.m files.
If you want to simulate the zincfinger concentrations, you need to do the following:
- In the file 'maincounter.m', set the parameters you want to simulate.
- In the Matlab terminal, type 'maincounter' in the command line.
- The computation takes about 2 seconds. Matlab then draws a plot where you can see the impact your chosen parameters have on the concentractions.
If you want to do the sensitivity analysis, do the following:
- In the file 'symsen_montecarlo.m' set the desired parameters to the variables para1 and para2 on line 62 and 63.
- Adapt the line 74 accordingly.
- In the Matlab terminal, type 'symsen_montecarlo' in the command line.
- The computation takes about 20 seconds. Matlab draws some plots and computes the correlation coefficients.
How does the concentration simulation work?
[http://www.tik.ee.ethz.ch/%7ebarkows/counter.zip Zincfinger concentrations gezippt]
maincounter.m
The main program is maincounter.m. First, the parameters are defined. Secondly, Matlab solves the differential equation (which are defined in ode_counter.m) using ode15s. Thirdly, a plot containing the input signal and all four concentrations is created. Last, Matlab calls the function jury.m which tells us whether the concentrations oscillates (which of course is what we want).
ode_counter.m
Here, the differential equations are defined. These are in words: The variation of a protein's concentration is equal to the product of activation and repression (because zincfingers act as roadblocks) minus a fixed part of the concentration itself (degradation).
act.m
Here, the equation we use for activation is defined.
rep.m
Here, the equation we use for reprimation is defined.
input_signal.m
The input signal is defined. You can choose between different input signals. The default input signal is a sinusoidal oscillation with 'breaks' (sections where the input remains zero between the sinusoidal peaks).
jury.m
Here, the function test_oscillation.m is called for every one of the four concentrations. jury.m returns one if any of the test_oscillation.m calls returned one (i.e. it suffices if one concentration is properly oscillating, because we can then choose that one to be the 'counting' concentration).
test_oscillation.m
Although the code looks quite complicating, what it does is pretty simple: It looks if the peaks (which of course have to occur with half the frequency of the input signal) are at least twice as big than the biggest point between the peaks (caused by noise).
What if I want to automatically test several sets of parameters?
You have set all parameters and done a simulation run. The result is negative (i.e. the concentrations don't oscillate correctly). You think you know which parameter you have to change to achieve a positive result, but you don't know how much you need to change it. There are two ways of finding that out:
- Change the parameter of interest by hand and do a simulation run. Look at the plot, and write down if that specific value works. Change the parameter again, do another simulation run, etc. If you want to test a large range of values (e.g. K from 1 to 1000 with stepsize 5), this approach is way too onerous (200 iterations). Instead, try the following:
- Use the function K_kd_iterate.m.
K_kd_iterate.m
Until now, only an iteration of K (zincfinger affinity) and kd (zincfinger degradation rate) has been implemented.
It works as follows: Create a vector with the values to be tested (of K and kd, respectively). Note that a higher K means a higher affinity (see the repression equation rep.m for details).
K_kd_iterate.m calls maincounter_iterate.m for every possible combination of K and kd. Be careful not to chose too large vectors. If you try 20 values for K and 20 values for kd, the computation takes a little more than ten (!) minutes.
maincounter_iterate.m
maincounter_iterate.m is the same as maincounter.m, only adapted to be used by K_kd_iterate.m (i.e. you can't set the parameters K and kd anymore).
How does the Sensitivity analysis work?
[http://people.ee.ethz.ch/~ulrichta/sensitivity.zip Sensitivity gezippt]
symsen_montecarlo.m
This file implements the Multiparametric Sensitivity Analysis (MPSA) based on the Monte Carlo Method.
Other files
The other three files (ode_counter.m, act.m, rep.m) are the same as in the zincfinger concentration simulation. They are used to define the differential equations that characterize our system.