Event Processing Device
From 2006.igem.org
Back to ETH Zurich main page.
Contents |
Purpose
In a nutshell, the Input module has 2 system boundaries, both of which are characterized by PoPS (Polymerase Per Second). Its purpose is to take a single input PoPS and output 2 different PoPS. One of the outputs should be high and the other low when S is high and vice versa when S is low.
PoPS_outn
^
¦
high ¦ PoPS_out2 _______ _________ PoPS_out1
¦ \ /
¦ \ /
¦ \ /
¦ \ /
¦ \ /
¦ X
¦ / \
¦ / \
¦ / \
¦ / \
low ¦ PoPS_out1 _______/ \_________ PoPS_out2
¦--------------------------------------------------> PoPS_in, t
0 1
Schematic
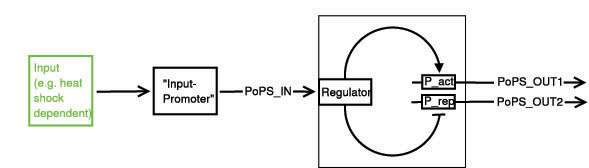
Below a preliminary parts-view of the module, i.e. encapsulation of biological specific implementations into a functional box with general PoPS interfaces.
With this design, any input can be used, as long as a promoter can be found that is either activated or suppressed by it. At the output, any kind of genes can be added which will be produced depending on the PoPS of that particular input.
legend: I: input signal that will bind to the promotor P_in, e.g. heat shock dependent etc. P_in: promotor that allows the signal of choice I to bind. PoPS_in: polymerase per second dependent on promotor and concentration of I Reg: regulation genes. there are different solutions possible, see below. r: repressing signal. highly dependent on Reg, P_r and of course speed and binding considerations a: activating signal. highly dependent on Reg, P_a and of course speed and binding considerations rep: repression, indicated by horizontal bar act: activation, indicated by arrow P_r: promotor region to be repressed and/or "roadblock" region. constitutively active P_a: promotor region to be activated and/or "roadblock" region. not constitutively active PoPS_out1: polymerase per second dependent on P_r and r. PoPS_out2: polymerase per second dependent on P_a and a. R_n: toggle switch gene(s)
Strategies & Evaluation
Suitable Strategies
As shown in the black box above, the core of the device is a regulator sub-module, for which different implementation strategies have been investigated.
We identified various solutions, the remaining ones being
- The λ-system with two variants
- The original anti-parallel version
- An engineered unidirectional version
- The Lux-system
Selected System
Due to limited time, the current availability of parts and the constraints due to the standard restriction sites available in the parts library, we decided to actually implement the unidirectional λ-system. As an input we will use IPTG for easy handling and debugging now, although the system is of course scalable for other types of inputs.
- Advantages of the λ-system
- Since the two promoters are regulated by the same protein-operator interactions, repression and activation should be symmetrical (which is crucial as modeling has shown).
- Parts are available from the registry package Registry 7.05.
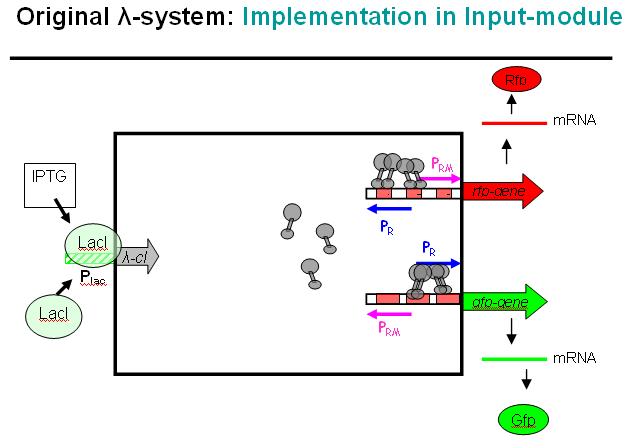
Basic Principle of Original λ-System
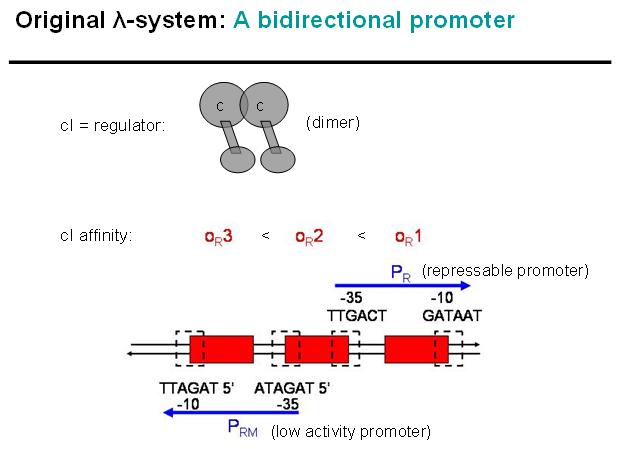
We decided on the λ-system to be implemented for now. The following schematics illustrate the basic concept with the natural λ-system (bidirectional), and below the the planned implementation of the unidirectional version in the INPUT-module.
Legend: cI is a dimer and regulates the activity of the two promoter regions, Pr and Prm, on the λ-system. Pr is constitutively active and is repressed when cI binds to the two operator regions it overlaps with, OR1 and OR2. Prm is not very active without cI, but has an unknown basal activity.
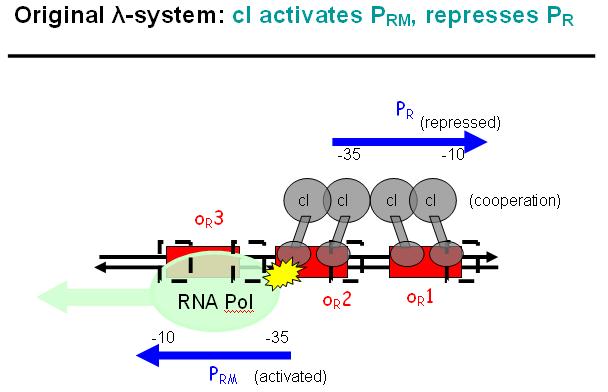
Legend: The binding affinity of the operator regions is different, i.e. OR1 > OR2 > OR3, such that cI first binds to OR1, then with cooperativity binds to OR2. As soon as cI is bound over the Pr promotor region, Pr will be repressed. However, the presence of cI at OR2 will make it possible for the RNApolymerase to bind to Prm and thus Prm will become active only in the presence of cI.
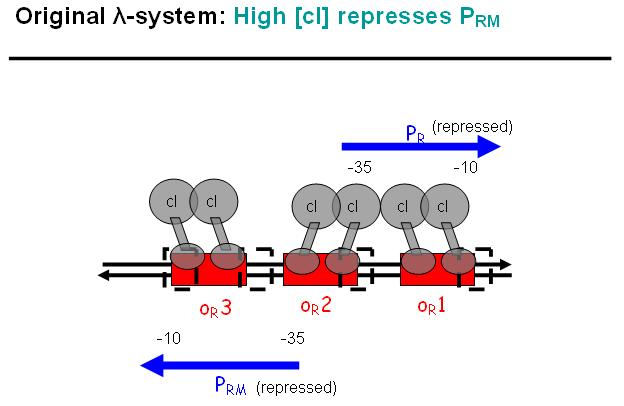
Legend: In a third stage, cI also binds to the OR3 region and thus repressed Prm again. However, in the modified system as it can be found in the registry, OR3 has been switched-off through mutation.
Basic Principle of Modified λ-System in INPUT-module
Legend: This schematic illustrates how the modified λ-system could be used as a regulator system in the INPUT-module. Here we would not use the original bidirectional system, but the unidirectional one that can be found in the registry, i.e. where Pr and Prm have been separated and OR3 deactivated over mutation.
At the input side, we could use the well known LacI-system. The LacI-protein would be present all the time, bind to the Lac-promoter (Plac) and thus block it. No cI (λ-cI) would be produced in this state. However, if IPTG is added to the system, it would remove the LacI-protein from the promoter and cI would be produced in this other state.
In absence of an input signal (i.e. no IPTG in this debugging system) no cI would be produced, Pr will be constitutively active and Prm would have low basal activity (which will be further repressed by the zinc-finger system, see NOR-module). In a debugging system as shown above, RFP would be produced and indicate this state with red fluorescence.
As soon as there is an input signal (i.e. IPTG activating the Lac-promoter), cI would be produced, thus repressing Pr and activating Prm. GFP would be produced, RFP production would cease. Thus, the red fluorescence would change to green fluorescence, if everything works as planned.
Actual Implementation
Full Sequence
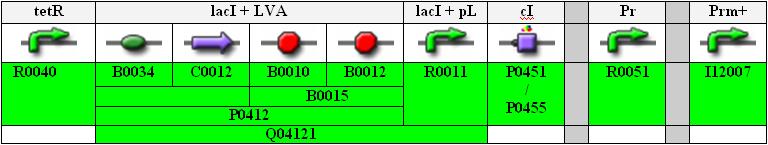
Below you find an illustration of the full sequence of the planned INPUT-module.
tetR is a constitutive promoter, thus lacI is continuously produced and repressing the lacI promoter.
Note that the Pr and Prm parts are separated (unidirectional, not bidirectional) and that the Prm+ part from the library is with a modified (non-functional) OR3 region.
Also, for the cI part we can either use the normal one, P0451, or the one with an additional degradation tag, P0455 (indicated in orange in the flow chart above).
Cloning Steps and Debugging
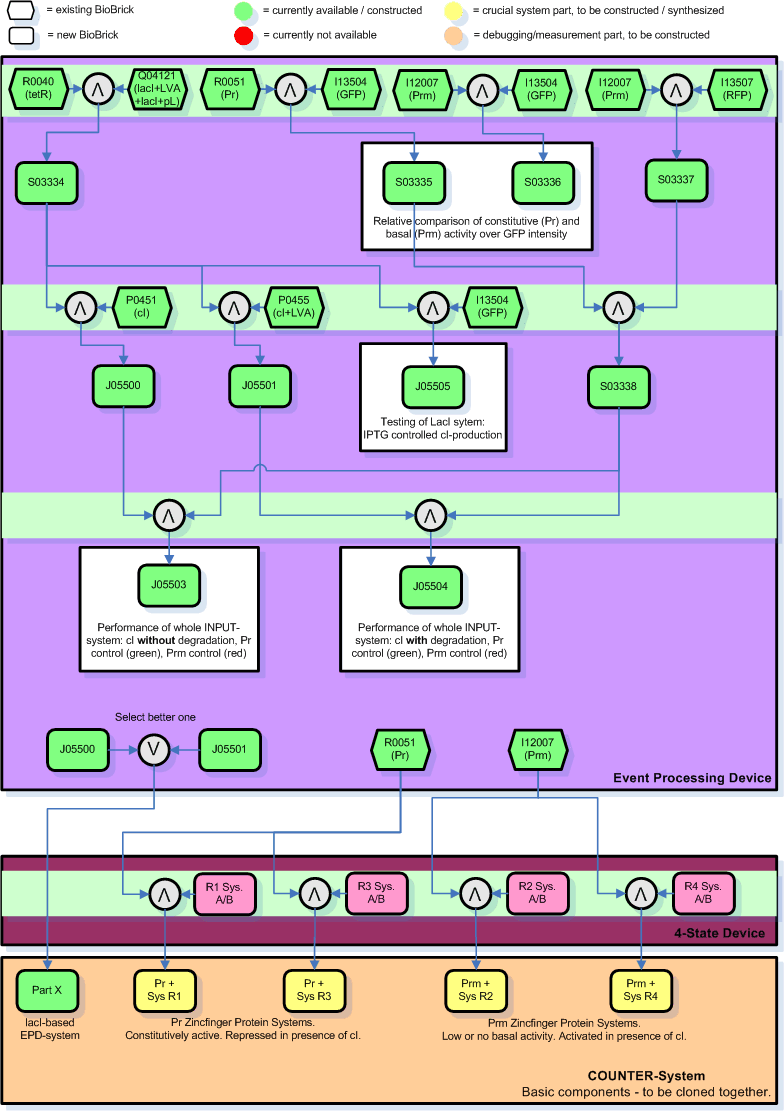
Below you find an illustration on how we plan to clone the input module. This graph will probably be revised soon according to inputs of the other members of the team who are more experienced with this stuff.
Legend:
- The number in the boxes correspond to the part number in the [http://parts.edu.mit registry].
- The yellow and orange parts are the intermediate combinations we will get as a result of cloning components together. These we will add to the registry.
- The orange parts are for debugging/testing purposes only. Note that we might chose to skip certain testing steps due to time constraints.
- There are 3 cloning sessions (indicated by light green horizontal bars) for the INPUT-module and its testing alone, each consisting of 2-4 individual cloning steps that can be done in parallel.
- The individual cloning steps are indicated by the symbol for a logical AND (arrow up in grey circle). When only the more suitable part has to be chosen to continue, a logical OR indicates that (arrow down in grey circle). The part to the right of the logical AND is attached downstream to the left part.
- The purpose of certain results, i.e. the testing, is commented in square boxes around the resulting part(s).
- The Zinc Hand parts with operator regions, here indicated by RX System A/B, developed and sythesized by the group working on the NOR-module are indicated in bright pink.
- For the final COUNTER-System will we will need to clone the Pr/Prm promoters at the beginning of each of the Zinc Hand systems and add the selected INPUT-module sequence (either intermediate part 4 or 5 - whichever is better) in front of everything. The final COUNTER-System will thus consist of 5 intermediate parts (here in the graph labeled as intermediates from X plus 11-14). It will take another 1 or 2 sessions to clone those components all together to a single system, depending on whether we want to put it on a single plasmid
Parts and Methods
The current status is that we will be able to clone the whole INPUT-module and its intermediate steps for debugging purposes over [http://parts2.mit.edu/r/parts/htdocs/Assembly/index.cgi standard assembly] with parts found in the package Registry 7.05.
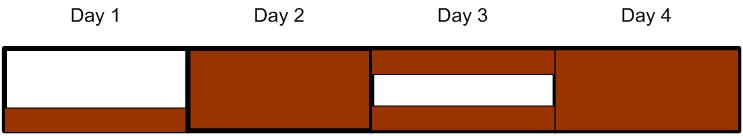
A single cloning session will typically take 4 days, if carried out by an experienced person and if no mistakes were made - or one week to play safe. The typical workload would roughly be something like the following diagram, where work is indicated in brown.
Cloning Schedule
Looks like we have 4 cloning sessions with each 4-5 days for the INPUT-module alone, which makes 16 to 20 days - if we want to do all the steps.
The actual cloning schedule can be found here.