Assembly
From 2006.igem.org
(→'''Parts to Make:''') |
|||
| (54 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | '''Parts to Make:''' | + | == '''Parts to Make:''' == |
HixL: TTCTGGAAAA CCAAGGTTTT TGATAA | HixL: TTCTGGAAAA CCAAGGTTTT TGATAA | ||
| Line 9: | Line 9: | ||
'''Backwards Parts''' | '''Backwards Parts''' | ||
| - | Because we wish to start with elements that are reversed (upside down pancakes), we need to think about how to get backwards parts. That is, parts that have the Prefix, then reversed element (promoter, term, CDS, RBS), then Suffix. Several possible ways: | + | Because we wish to start with elements that are reversed (upside down pancakes), we need to think about how to get backwards parts. That is, parts that have the Prefix, then reversed element (promoter, term, CDS, RBS), then Suffix. |
| + | |||
| + | A good way to make a backwards part is to PCR amplify the part from a plasmid with primers that include a XbaI site on the 3' end of the part and a SpeI on the 5' end (and four extra bases). Then the PCR product can be digested with XbaI and SpeI and directionally cloned into a vector. | ||
| + | |||
| + | For the reverse orientation of the Ara-BAD promoter (BBa_I13453), we designed these primers: | ||
| + | |||
| + | pARA-Spe-F | ||
| + | |||
| + | 5’ GCAT ACTAGT ACATTGATTATTTGCACGGC 3’ | ||
| + | |||
| + | pARA-Xba-R | ||
| + | |||
| + | 5’ GCAT TCTAGA GCTAGCCCAAAAAAACGGTA 3’ | ||
| + | |||
| + | Alternative plan for making pARA reverse uses these oligos: | ||
| + | |||
| + | pBAD_rev_Top | ||
| + | |||
| + | 5’ GCAT ACTAGT ACATTGATTATTTGCACGGCGTCACACTTTGCTATGCCATAGCATTTTTATCCATAAGATTAGCGGATCC 3’ | ||
| + | |||
| + | pBAD_rev_Bottom | ||
| + | |||
| + | 5’ GCAT TCTAGA GCTAGCCCAAAAAAACGGTATGGAGAAACAGTAGAGAGTTGCGATAAAAAGCGTCAGGTAGGATCCGCTA 3’ | ||
| + | |||
| + | For the reverse orientation of the RBS - BBa_B0030 Plate DNA-1, Spot 3G, pSB1A2, we designed these primers: | ||
| + | |||
| + | RBS-back-Spe-F | ||
| + | |||
| + | 5’ GCAT ACTAGT ATTAAAGAGGAGAAA 3’ | ||
| + | |||
| + | RBS-back-R | ||
| + | |||
| + | 5’ GCAT TCTAGA TTTCTCCTCTTTAAT 3’ | ||
| + | |||
| + | For the reverse orientation of the double forward terminator, BBa_B0015 Plate DNA-1, Spot 1I, pSB1AK3, we designed these primers: | ||
| + | |||
| + | TT-back-Spe-F | ||
| + | |||
| + | 5’ GCAT ACTAGT CCAGGCATCAAATAAAACGA | ||
| + | |||
| + | TT-back-R | ||
| + | |||
| + | 5’ GCAT TCTAGA TATAAACGCAGAAAGGCCCA | ||
| + | |||
| + | Several other possible ways: | ||
# Some parts are backwards in the registry - terminators, for eg. | # Some parts are backwards in the registry - terminators, for eg. | ||
# Parts that are not long could be synthesized backwards. | # Parts that are not long could be synthesized backwards. | ||
# If one pancake can be flipped, reversed parts can be made. This may actually be a good contribution, since it could be applied to reverse any part between two hix sites. | # If one pancake can be flipped, reversed parts can be made. This may actually be a good contribution, since it could be applied to reverse any part between two hix sites. | ||
| + | # Can cut out a part with NotI, then religate in both orientations. | ||
Examples | Examples | ||
| Line 19: | Line 64: | ||
BBa_B0022 Terminator (Reverse B0012) Length: 84 Base Pairs Including BioBrick Ends | BBa_B0022 Terminator (Reverse B0012) Length: 84 Base Pairs Including BioBrick Ends | ||
| - | + | BBa_P1004 is ampR resistance cassette in reverse orientation Length: 984 Base Pairs Including BioBrick Ends (In planning stages only) | |
Bba_B0034 is a RBS with the sequence: | Bba_B0034 is a RBS with the sequence: | ||
| - | gaattcgcggccgcttctagag ''' | + | gaattcgcggccgcttctagag '''attaaagaggagaaa''' tactagtagcggccgctgcag |
It could be redesigned with the RBS reversed: | It could be redesigned with the RBS reversed: | ||
| - | gaattcgcggccgcttctagag ''' | + | gaattcgcggccgcttctagag '''tttctcctctttaat''' tactagtagcggccgctgcag |
| + | Oligos for reverse RBS that produce EcoRI and SpeI sticky ends: | ||
| - | + | 5’ AATTC GCGGCCGC T TCTAGA G TTTCTCCTCTTTAAT T A 3’ | |
| - | + | 5’ CTAGT A ATTAAAGAGGAGAAA C TCTAGA A GCGGCCGC G 3’ | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | Oligos for reverse pBAD promoter that produce XbaI and SpeI sites with 4 extra bp on ends and 10 bp overlap in middle: | |
| - | + | ||
| - | + | ||
| - | + | pBAD_rev_Top | |
| - | + | 5' - GCAT ACTAGTA CATTGATTATTTGCACGGCGTCACACTTTGCTATGCCATAGCATTTTTATCCATAAGAT TAGCGGATCC - 3' | |
| - | + | pBAD_rev_Bottom | |
| - | + | 5' - GCAT TCTAGA GCTAGCCCAAAAAAACGGTATGGAGAAACAGTAGAGAGTTGCGATAAAAAGCGTCAGGTA GGATCCGCTA - 3' | |
| - | + | ||
| - | + | ||
| - | |||
| - | + | Here is a proposed strategy for making backwards parts flanked by hix sites: | |
| - | + | [[Image:Backwards.jpg]] | |
== '''hix BioBricks''' == | == '''hix BioBricks''' == | ||
| - | |||
'''E - N - X - hix - S - N - P''' | '''E - N - X - hix - S - N - P''' | ||
| - | Does there have to be a T between NotI and XbaI and an A between SpeI and NotI? All the parts we looked at | + | Does there have to be a T between NotI and XbaI and an A between SpeI and NotI? All the parts we have looked at so far have this. |
HixL | HixL | ||
| - | + | GAATTC GCGGCCGC T TCTAGA TTCTTGAAAACCAAGGTTTTTGATAA ACTAGT A GCGGCCGC CTGCAG | |
HixR | HixR | ||
| - | + | GAATTC GCGGCCGC T TCTAGA TTATCAAAAACCTTCCAAAAGGAAAA ACTAGT A GCGGCCGC CTGCAG | |
HixC | HixC | ||
| - | + | GAATTC GCGGCCGC T TCTAGA TTATCAAAAACCATGGTTTTTGATAA ACTAGT A GCGGCCGC CTGCAG | |
'''Oligos to order''' | '''Oligos to order''' | ||
| Line 80: | Line 117: | ||
HixL_top | HixL_top | ||
| - | + | AATTCGCGGCCGCTTCTAGATTCTTGAAAACCAAGGTTTTTGATAAACTAGTAGCGGCCGCCTGCA | |
HixL_bottom | HixL_bottom | ||
| - | + | GGCGGCCGCTACTAGTTTATCAAAAACCTTGGTTTTCAAGAATCTAGAAGCGGCCGCG | |
HixR_top | HixR_top | ||
| - | + | AATTCGCGGCCGCTTCTAGATTATCAAAAACCTTCCAAAAGGAAAAACTAGTAGCGGCCGCCTGCA | |
HixR_bottom | HixR_bottom | ||
| - | + | GGCGGCCGCTACTAGTTTTTCCTTTTGGAAGGTTTTTGATAATCTAGAAGCGGCCGCG | |
HixC_top | HixC_top | ||
| - | + | AATTCGCGGCCGCTTCTAGATTATCAAAAACCATGGTTTTTGATAAACTAGTAGCGGCCGCCTGCA | |
HixC_bottom | HixC_bottom | ||
| - | + | GGCGGCCGCTACTAGTTTATCAAAAACCATGGTTTTTGATAATCTAGAAGCGGCCGCG | |
| + | == '''Building''' == | ||
| - | ''' | + | '''Biobricking in one hix site:''' |
| - | [[Image: | + | [[Image:Hix ligation11.JPG]] |
| + | |||
| + | '''Example of a construct for one flip (terminator, although CDS would be better):''' | ||
[[Image:HIX_Final.JPG]] | [[Image:HIX_Final.JPG]] | ||
| + | |||
| + | '''Biobricking hix sites onto parts:''' | ||
| + | |||
| + | [[Image:Pan1.JPG]] | ||
| + | |||
| + | '''Assembly of One Pancake Test Construct:''' | ||
| + | |||
| + | |||
| + | [[Image:Pan2.JPG]] | ||
| + | |||
| + | '''Assembly of Two Pancake Test Construct would be next. Probably want two genes.''' | ||
| + | |||
| + | '''Assembly of Multiple Pancake Test Constructs:''' | ||
| + | |||
| + | [[Image:Pan3.JPG]] | ||
| + | |||
| + | == '''First Transformations:''' == | ||
| + | |||
| + | >1. Terminator T1 Bba_B0010 DNA-2 3P< | ||
| + | |||
| + | 2. Promoter Pbad BBa_I13453 | ||
| + | |||
| + | >3. Promoter w/RBS (TetR Repressed) BBa_J13002 DNA-2 3F < | ||
| + | |||
| + | 4. Reverse Amp^r BBa_P1004 (Oops, planning) | ||
| + | |||
| + | 5. Reverse CmR BBa_P1009 789 bp (Oops, planning) | ||
| + | |||
| + | >6. Double Forward Terminator BBa_B0015 DNA-1 1I < | ||
| + | |||
| + | >7. Tet^r Bba_P1001 DNA-2 23F < | ||
| + | |||
| + | == '''Questions:''' == | ||
| + | |||
| + | Will the TetR repressed promoter in BBa_J13002 be always on in JM109 cells? | ||
| + | |||
| + | Do we want to be able to turn on a Start Promoter(i.e. LacP using IPTG) on/off so recombination occurs 1st; and then use a different inducible promoter to turn on transcription? | ||
Latest revision as of 18:08, 20 July 2006
Contents |
Parts to Make:
HixL: TTCTGGAAAA CCAAGGTTTT TGATAA
HixR: TTATCAAAAA CCTTCCAAAA GGAAAA
HixC: TTATCAAAAA CCATGGTTTT TGATAA
Backwards Parts
Because we wish to start with elements that are reversed (upside down pancakes), we need to think about how to get backwards parts. That is, parts that have the Prefix, then reversed element (promoter, term, CDS, RBS), then Suffix.
A good way to make a backwards part is to PCR amplify the part from a plasmid with primers that include a XbaI site on the 3' end of the part and a SpeI on the 5' end (and four extra bases). Then the PCR product can be digested with XbaI and SpeI and directionally cloned into a vector.
For the reverse orientation of the Ara-BAD promoter (BBa_I13453), we designed these primers:
pARA-Spe-F
5’ GCAT ACTAGT ACATTGATTATTTGCACGGC 3’
pARA-Xba-R
5’ GCAT TCTAGA GCTAGCCCAAAAAAACGGTA 3’
Alternative plan for making pARA reverse uses these oligos:
pBAD_rev_Top
5’ GCAT ACTAGT ACATTGATTATTTGCACGGCGTCACACTTTGCTATGCCATAGCATTTTTATCCATAAGATTAGCGGATCC 3’
pBAD_rev_Bottom
5’ GCAT TCTAGA GCTAGCCCAAAAAAACGGTATGGAGAAACAGTAGAGAGTTGCGATAAAAAGCGTCAGGTAGGATCCGCTA 3’
For the reverse orientation of the RBS - BBa_B0030 Plate DNA-1, Spot 3G, pSB1A2, we designed these primers:
RBS-back-Spe-F
5’ GCAT ACTAGT ATTAAAGAGGAGAAA 3’
RBS-back-R
5’ GCAT TCTAGA TTTCTCCTCTTTAAT 3’
For the reverse orientation of the double forward terminator, BBa_B0015 Plate DNA-1, Spot 1I, pSB1AK3, we designed these primers:
TT-back-Spe-F
5’ GCAT ACTAGT CCAGGCATCAAATAAAACGA
TT-back-R
5’ GCAT TCTAGA TATAAACGCAGAAAGGCCCA
Several other possible ways:
- Some parts are backwards in the registry - terminators, for eg.
- Parts that are not long could be synthesized backwards.
- If one pancake can be flipped, reversed parts can be made. This may actually be a good contribution, since it could be applied to reverse any part between two hix sites.
- Can cut out a part with NotI, then religate in both orientations.
Examples
BBa_B0022 Terminator (Reverse B0012) Length: 84 Base Pairs Including BioBrick Ends
BBa_P1004 is ampR resistance cassette in reverse orientation Length: 984 Base Pairs Including BioBrick Ends (In planning stages only)
Bba_B0034 is a RBS with the sequence:
gaattcgcggccgcttctagag attaaagaggagaaa tactagtagcggccgctgcag
It could be redesigned with the RBS reversed:
gaattcgcggccgcttctagag tttctcctctttaat tactagtagcggccgctgcag
Oligos for reverse RBS that produce EcoRI and SpeI sticky ends:
5’ AATTC GCGGCCGC T TCTAGA G TTTCTCCTCTTTAAT T A 3’
5’ CTAGT A ATTAAAGAGGAGAAA C TCTAGA A GCGGCCGC G 3’
Oligos for reverse pBAD promoter that produce XbaI and SpeI sites with 4 extra bp on ends and 10 bp overlap in middle:
pBAD_rev_Top
5' - GCAT ACTAGTA CATTGATTATTTGCACGGCGTCACACTTTGCTATGCCATAGCATTTTTATCCATAAGAT TAGCGGATCC - 3'
pBAD_rev_Bottom
5' - GCAT TCTAGA GCTAGCCCAAAAAAACGGTATGGAGAAACAGTAGAGAGTTGCGATAAAAAGCGTCAGGTA GGATCCGCTA - 3'
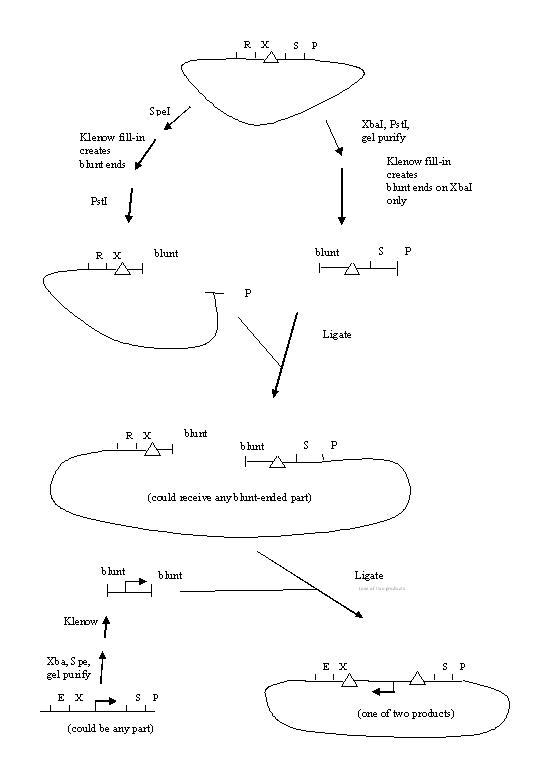
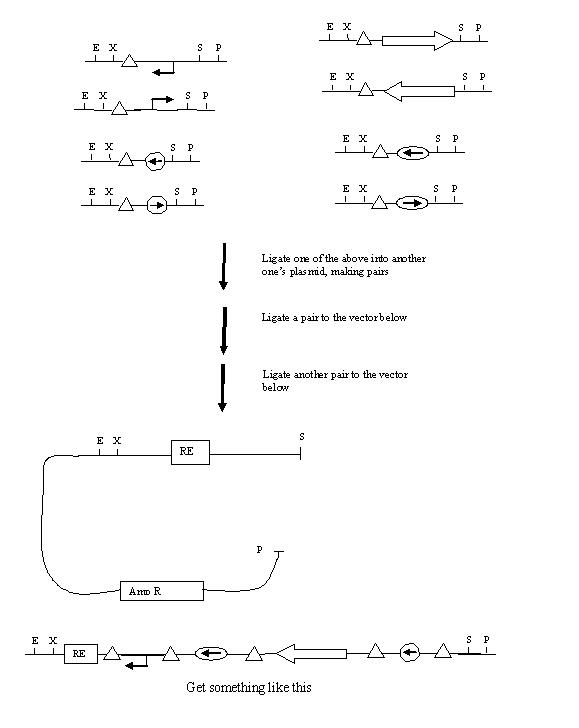
Here is a proposed strategy for making backwards parts flanked by hix sites:
hix BioBricks
E - N - X - hix - S - N - P
Does there have to be a T between NotI and XbaI and an A between SpeI and NotI? All the parts we have looked at so far have this.
HixL
GAATTC GCGGCCGC T TCTAGA TTCTTGAAAACCAAGGTTTTTGATAA ACTAGT A GCGGCCGC CTGCAG
HixR
GAATTC GCGGCCGC T TCTAGA TTATCAAAAACCTTCCAAAAGGAAAA ACTAGT A GCGGCCGC CTGCAG
HixC
GAATTC GCGGCCGC T TCTAGA TTATCAAAAACCATGGTTTTTGATAA ACTAGT A GCGGCCGC CTGCAG
Oligos to order
HixL_top
AATTCGCGGCCGCTTCTAGATTCTTGAAAACCAAGGTTTTTGATAAACTAGTAGCGGCCGCCTGCA
HixL_bottom
GGCGGCCGCTACTAGTTTATCAAAAACCTTGGTTTTCAAGAATCTAGAAGCGGCCGCG
HixR_top
AATTCGCGGCCGCTTCTAGATTATCAAAAACCTTCCAAAAGGAAAAACTAGTAGCGGCCGCCTGCA
HixR_bottom
GGCGGCCGCTACTAGTTTTTCCTTTTGGAAGGTTTTTGATAATCTAGAAGCGGCCGCG
HixC_top
AATTCGCGGCCGCTTCTAGATTATCAAAAACCATGGTTTTTGATAAACTAGTAGCGGCCGCCTGCA
HixC_bottom
GGCGGCCGCTACTAGTTTATCAAAAACCATGGTTTTTGATAATCTAGAAGCGGCCGCG
Building
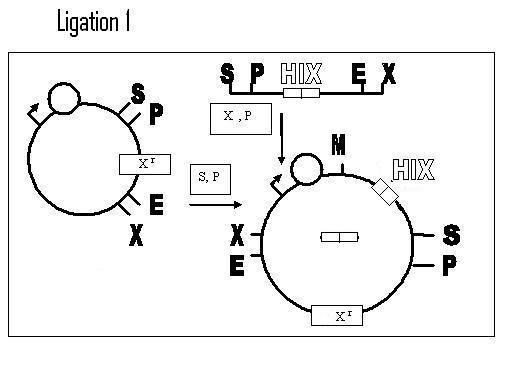
Biobricking in one hix site:
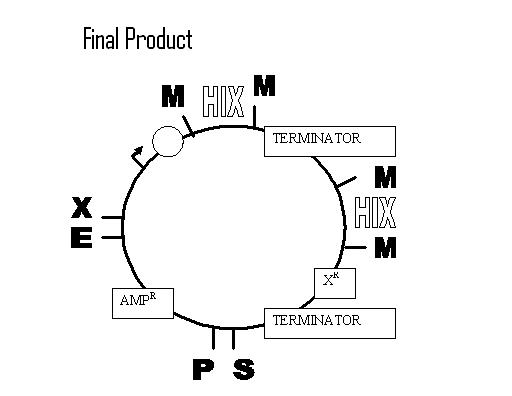
Example of a construct for one flip (terminator, although CDS would be better):
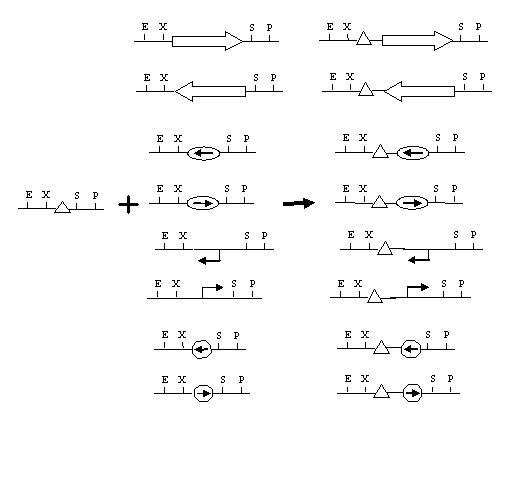
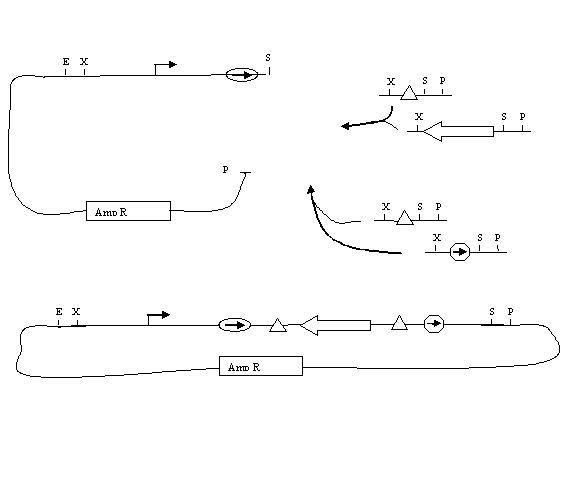
Biobricking hix sites onto parts:
Assembly of One Pancake Test Construct:
Assembly of Two Pancake Test Construct would be next. Probably want two genes.
Assembly of Multiple Pancake Test Constructs:
First Transformations:
>1. Terminator T1 Bba_B0010 DNA-2 3P<
2. Promoter Pbad BBa_I13453
>3. Promoter w/RBS (TetR Repressed) BBa_J13002 DNA-2 3F <
4. Reverse Amp^r BBa_P1004 (Oops, planning)
5. Reverse CmR BBa_P1009 789 bp (Oops, planning)
>6. Double Forward Terminator BBa_B0015 DNA-1 1I <
>7. Tet^r Bba_P1001 DNA-2 23F <
Questions:
Will the TetR repressed promoter in BBa_J13002 be always on in JM109 cells?
Do we want to be able to turn on a Start Promoter(i.e. LacP using IPTG) on/off so recombination occurs 1st; and then use a different inducible promoter to turn on transcription?