University of California San Francisco 2006
From 2006.igem.org
(→Aspirations, Dreams...) |
(→The Orthogonal Interaction and Receptor Specificity) |
||
| Line 25: | Line 25: | ||
== The Orthogonal Interaction and Receptor Specificity == | == The Orthogonal Interaction and Receptor Specificity == | ||
To selectively discriminate between the chemoreceptors, we will re-engineer the signaling protein ''CheW'' (which does not differentiate between receptors in its native state), to recognize only one receptor. This will require the development of an orthogonal interaction - that is two interacting partners that have no cross-talk between orhtogonal partners. | To selectively discriminate between the chemoreceptors, we will re-engineer the signaling protein ''CheW'' (which does not differentiate between receptors in its native state), to recognize only one receptor. This will require the development of an orthogonal interaction - that is two interacting partners that have no cross-talk between orhtogonal partners. | ||
| - | + | [[Image:cartoon.jpg|thumb|left|250px|''An orthogonal interaction for all seasons'']] | |
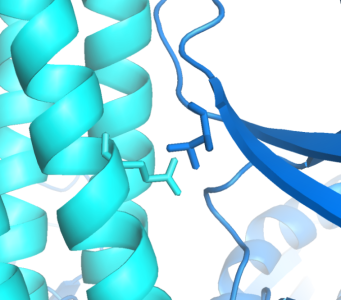
Implementing an orthogonal interaction typicaly requires detailed knowledge of the interface. While no structure for the ''CheW-Tar'' complex yet exists, their unbound structures have been characterized. By computationally docking them together, we can observe that residues identified by the Parkinson group as to having destabilizing/recovery properties for binding do indeed make contact in complex. This interface can be further redesigned using computational approaches. | Implementing an orthogonal interaction typicaly requires detailed knowledge of the interface. While no structure for the ''CheW-Tar'' complex yet exists, their unbound structures have been characterized. By computationally docking them together, we can observe that residues identified by the Parkinson group as to having destabilizing/recovery properties for binding do indeed make contact in complex. This interface can be further redesigned using computational approaches. | ||
| + | [[Image:complex1.jpg|thumb|left|250px|''Docked and loaded'']] | ||
| + | Now we also need two receptors which are responsive to different attractants. | ||
Much work is available in the literature regarding the changing of chemoreceptor specificity. Some of the options include: | Much work is available in the literature regarding the changing of chemoreceptor specificity. Some of the options include: | ||
Revision as of 21:59, 30 October 2006
Contents |
The University of California at San Francisco 2006 iGEM Team
The international Genetically Engineered Machine Competition is an opportunity for emerging young scientists of all disciplines to tinker with the proverbial Rube Goldberg machine of life. This is the UCSF team's third year in the program, and to celebrate our three years, we had a members-only contest to develop a team logo. More information on the logo is forthcoming.
Team Members
- Ala Trusina
- Patrick Visperas
- Kevin Shay
- Matt Eames
Faculty Advisors
- [http://www.voigtlab.ucsf.edu/ Chris Voigt]
- [http://kortemmelab.ucsf.edu/ Tanja Kortemme]
Aspirations, Dreams...
What if we could control chemotaxis? Could we put bacteria on plate and steer them through a maze? Would it be like operating a remote-controlled car?
These were the questions that inspired team UCSF to tinker with bacterial chemotaxis. Naturally, bacteria don't have the same electric components that your remote-controlled buggy does, but there are parts we can harness to control motility. Escherichia coli, for example, already possess five different chemoreceptors which respond to different attractants and repellants. If we were to place the bugs in between two separate chemicals, we might be able to control the direction of swimming by manipulating which receptor's signal we interpret.
The UCSF iGEM team has endeavored to control chemotaxis by selectively expressing CheW mutants which will participate in an orthogonal interaction with the methyl-accepting chemotaxis protein Tar. The project will be broken down into three major components:
- Design and construction of the orthogonal interaction: An existing orthogonal interaction, to be used as a starting point prior to computational redesign, was discovered by [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=1860813&query_hl=2&itool=pubmed_docsum Liu and Parkinson] in 1991.
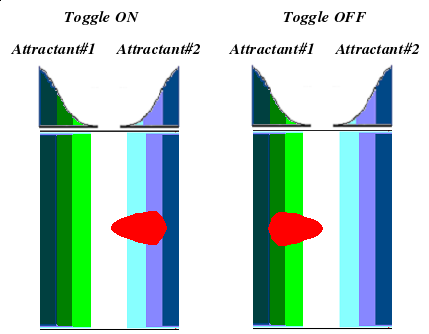
- Implementation of the genetic switch: This will allow us to toggle between the selective expression of either CheW.
- Validation of chemotactic properties through specially developed assay: originally developed by the good folks at the [http://www.physics.upenn.edu/facultyinfo/goulian.html Goulian Lab] in 2006.
The Orthogonal Interaction and Receptor Specificity
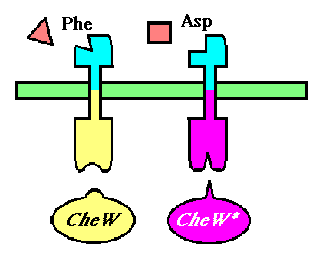
To selectively discriminate between the chemoreceptors, we will re-engineer the signaling protein CheW (which does not differentiate between receptors in its native state), to recognize only one receptor. This will require the development of an orthogonal interaction - that is two interacting partners that have no cross-talk between orhtogonal partners.
Implementing an orthogonal interaction typicaly requires detailed knowledge of the interface. While no structure for the CheW-Tar complex yet exists, their unbound structures have been characterized. By computationally docking them together, we can observe that residues identified by the Parkinson group as to having destabilizing/recovery properties for binding do indeed make contact in complex. This interface can be further redesigned using computational approaches.
Now we also need two receptors which are responsive to different attractants.
Much work is available in the literature regarding the changing of chemoreceptor specificity. Some of the options include:
- Directed evolution
- Computational redesign
- Chimeric construction (perisplasmic and cytoplasmic domains of the different receptors can be swapped)
We will be using a mutant version of the E. coli Tar (asparate-sensitive) receptor that has been evolved to respond to phenylalanine, a natural chemorepellant. The responsiveness of this mutant has been characterized and was found to have comparable chemotactic capabilities to that of wild-type Tar.
Chassis
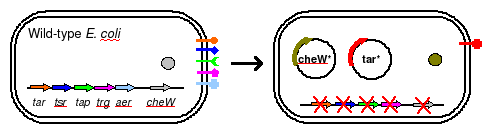
To eliminate interference from other naturally occuring receptors in E. coli, a strain with all receptors (Tar, Tsr, Tap, Trg, Aer) knocked out, provided by the Parkinson Lab, will be used in this work. The UCSF iGEM deleted CheW from this strain so that all CheW might be expression off the plasmid.
The plasmid will posses both constituitively active Tar varients, and will incorporate selective CheW control by means of a genetic switch.
Thanks
- [http://www.chemeng.uiuc.edu/Faculty/rao.html Chris Rao]
- [http://www.biology.utah.edu/faculty2.php?inum=6 Parkinson Lab]
- [http://www.physics.upenn.edu/facultyinfo/goulian.html Goulian Lab]
Links
- [http://www.ucsf.edu UCSF]