4-State Device
From 2006.igem.org
Back to the ETH Zurich main page.
Contents |
Organisation
Group Members
- Alexander Roth(coordinator), Urs A. Müller, Simon Barkow, Tamara Ulrich, Robin Künzler, Herve, Dominic, Christophe ...
Meetings
2005.08.17, Wednesday, 15:30 @ Alex' bureau (CAB F62) Summary 2005.08.18, Thursday, 13:15 @ Alex' bureau (CAB F62) 2005.08.22, Monday, 13:45 @ Alex' bureau (CAB F62) Summary 050822 2005.08.25, Thursday 15:30 @ Alex' bureau (CAB F62) 2005.08.29, Monday 14:00 @ Alex' bureau (CAB F62) 2005.08.30, Tuesday 16:00 @ Alex' bureau (CAB F62) 2005.09.06, Tuesday 08:00 @ Alex' bureau (CAB F62)
Description of NOR module
This module behaves like a NOR gate. It has two inputs and one output. The output is high only when neither input 1 nor input 2 is high.
NOR-module Schematic
Below a preliminary parts-view of the module with PoPS interfaces (i/o).
---------------------------------
| ------------\ |
PoPS_in1 ---->| | Repressor1 | -------- |
| ------------/ | |
| = |
| --------|------> PoPS_out
| = |
| ------------\ | |
PoPS_in2 ---->| | Repressor2 | -------- |
| ------------/ |
---------------------------------
Design
Zinc Fingers
A zinc finger [1] is a protein domain that binds to double stranded DNA. The idea is to use a zinc finger protein (ZFP) as a repressor by putting a ZFP binding site upstream of the coding region and thereby preventing RNA polymerase to transcribe the gene. A ZFP consistists of one or several Zinc Finger Domains that binds to three base pairs each. Many domain-DNA interaction for a triplet of base pairs have been described, therefore making it possible to to construct artificial transcription factors by combining domains.
Zinc Finger Proteins
In total for the counter we will need four ZFPs. We have two different designs of the ZFPs.
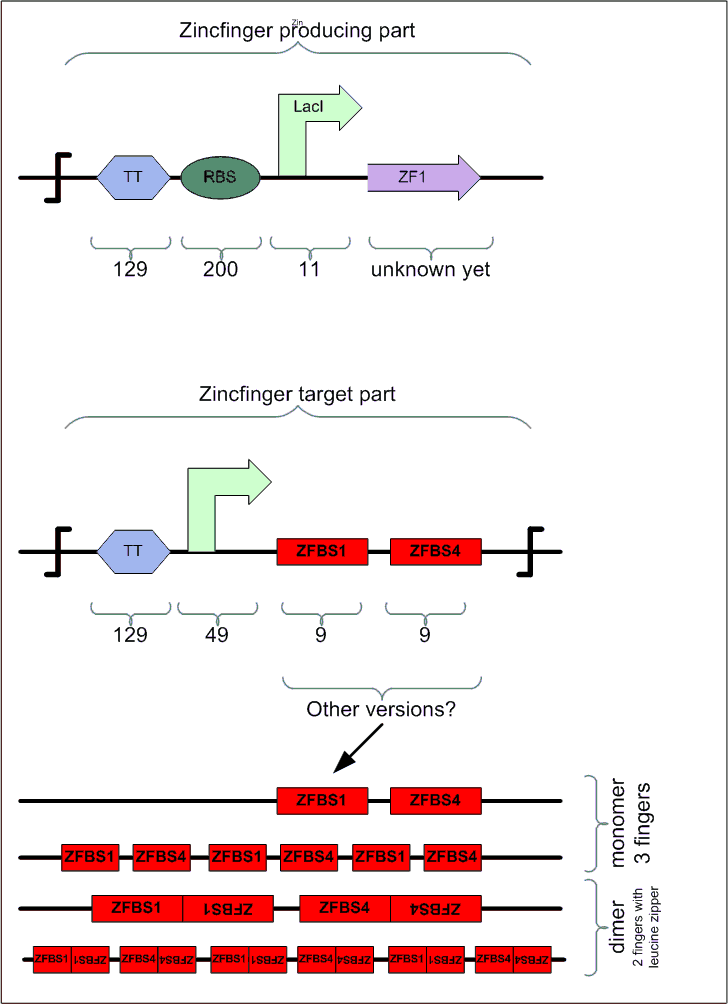
1. Three domains to recognize nine base pairs.
2. two zinc finger domains and a leucine zipper domain. The leucine zipper will be cause the protein to home-dimerize and hence it will recognize 12 base pairs.
Zinc Finger Protein Binding Sites
Since we are going to synthesize parts of our module, we have the possibility to design our own ZFP binding sites, as well as our ZFPs. We have to different candidates for placing the binding sites.
1. Binding sites in the promoter. This would prevent the polymerase from binding to the promotor.
2. Binding sites directly after the start of transcription and before the ribosome binding site. This alternative is quite attractive, since it would allow for a high degree of modularity. In theory the ZFP would act as an extra "roadblock-operator" and any promotor could be used in front of the protein.
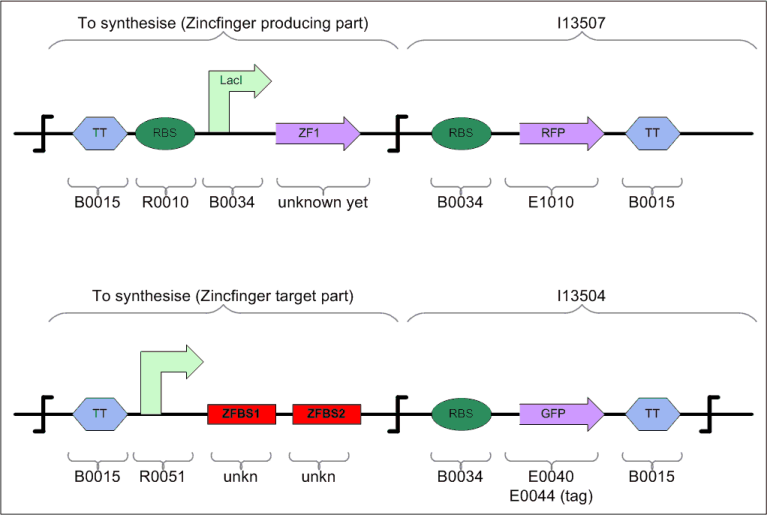
Tester for Zinc Finger
To test whether our assumptions about using multiple zinc finger proteins (ZFP) as repressors (i.e. roadblocks) will actually work, we will build a tester/debugging device in parallel with the counter .
System Variants
Currently, we have two systems, and we hope that at least one of them is going to work.
The first system, cryptically labelled "3dZFp-R" below, consists of 4 proteins with 4 operator regions. Each protein has four domains: a) 3 zink fingers that bind to specific sites on the helical DNA, and b) one repression (R) domain. We call the whole system (i.e. a+b) "Zinc Hand" (ZH) to distinguish it clearly from the individual zinc finger domains.
In the second system, labelled "2dZFp-LZ-R" (LZ = leucine zipper), each protein has also 4 domains: 2 zink fingers, 1 leucine zipper (LZ) and 1 repression (R) domain.
The operator regions in both systems are made of 2 binding regions, each of which lets a specific Zinc Hand bind to it and thus each consisting of the corresponding 3 codons that match the 3 zinc fingers in that Zinc Hand. The operator regions (i.e. the "roadblocks" that will prevent the binding of the RNApolymerase) form a BioBrick that should be inserted between the promoter region and the ribosome binding site in order to keep the design modular. Since we always use 2 Zinc Hands to block a promoter site (i.e. the promoters Pr and Prm of the lambda-system) and since we have 4 such promoter sites (i.e. 2xPr and 2xPrm), we need to have 4 operators = 4 permutations of Zinc Hand pairs.
ZHR stands for Zink Hands Regulatory proteins, and ZHO for Zink Hands Operators. The +1/+10 indication reflects the starting position of the 2 binding regions on the DNA. In both case, it between the promoter region (<0) and the ribosome binding site (around +20). There has been some discussion about having the operator around 0 (-9/+1)
| Type | Size (bp) per Unit | . | . | . | . | TOTAL SYSTEM SIZE (bp) | |
|---|---|---|---|---|---|---|---|
| 3dZFp-R | Repressor (protein) | 400 | ZHR1 | ZHR2 | ZHR3 | ZHR4 | 1600 |
| . | Operator +1/+10 (target sequence) | 30 | ZHO1.1 | ZHO2.1 | ZHO3.1 | ZHO4.1 | 120 |
| 2dZFp-LZ-R | Repressor (protein) | 400 | ZHR5 | ZHR6 | ZHR7 | ZHR8 | 1600 |
| . | Operator +1/+10 (target sequence) | 30 | ZHO5 | ZHO6 | ZHO7 | ZHO8 | 120 |
| TOTAL | . | . | . | . | . | . | 3440 |
Repression
Note that in both cases, it is still under discussion as whether the repression domain should really be included or not. If it turns out that repression has been used in all the literature we (well, actually Hervé) can find, then we would be better off to play safe and include them:
- Beerli PNAS 1998: they fused ZF to KRAB repressor and it has a stronger effect compare to ZF alone (see p14632 graph A)
- Beerli Nat Biotech review feb2002 : if you read the complete paragraph on gene repression (p 132), it gives strong evidence that we should fuse the ZF to a repressor (apparently at the N-Term of the ZF) (either KRAB or SID repressor). It is stated that polymerase blockade through ZF only is not very efficient.
However, the discussion is ongoing: As Alex pointed out we can not be sure that the 45 amino acid long KRAB domain will work, if it don't exists in prokaryotes. KRAB domains have a very specific interaction with a co-repressor molecule. It might be better to find something already known to work in bacteria or skip the repressor approach altogether. An alternative could be a negative transcription domain fused to the ZFP.
Degradation
We considered adding a variant with degradation tag, since the system might be slow otherwise. However, due to the fact that we have limited resources - in terms of money, manpower, and time - and the fact that the degradation is decoupled and it will be clearly visible if the system turns out to be too slow, we decided against it for now.
Additional Comments
We changed our strategy a couple of times in heated debates. There is quite obviously a trade-off between probability of success through redundancy, i.e. trying multiple approaches in parallel, and limited resources, i.e. financial, manpower, and time. Thus we decided against certain additional variants and cloning steps since we simple won't have the time for it. Another important aspect is the overall goal of keeping the design modular - one of the most important aspects of the this contest. The modification of the promoter regions of the lambda-system, i.e. Pr and Prm, although a valid approach will probably be dropped, since it is too much work and cost and no longer modular (as opposed to having the operators after the promoter).
Assembly
- Cut each system into BioBricks Pieces using different restriction enzymes.
- Assemble in parallel for each genes 1-4 in two variants: promoter+operator, rbs+gene
- Assemble the two constructs
- ...
(please complete the above and add the test/debugging steps)
References
Kim & Wang (??)
Park et al. 2005 (Activation of transcription)
Chou et al. 1998
Beerli00 (Linker)
Segal99 ("GNN-paper")
Dreier01 ("ANN-paper")
Dreier05 ("CNN-paper")
Yang95 (Kinetics!!)
Klug05 (Minireview)
Newman03 ("Leucine Zippers")