Imperial College 2006
From 2006.igem.org

|
|
|
Main Project
Secondary Projects
| [http://openwetware.org/wiki/IGEM:IMPERIAL/2006/project/popsblocker PoPs Blocker] | [http://openwetware.org/wiki/IGEM:IMPERIAL/2006/project/Bio_elec_interface Biological to Electrical Interface] |
|---|---|
|
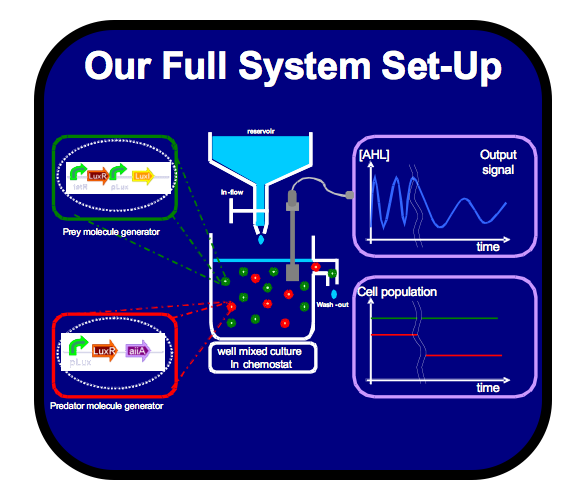
As a method of controlling the activation of the positive-feedback loop in our predator-prey based oscillator, we successfully created this part, which can be used as a general Pops Blocker:
|
We also worked on a Biosensor for measuring AHL concentrations in order to establish a Biological to Electrical Interface this summer.
|
Our Contributions to the Registry
| Part Logo | Description | Link to registry | Built | Tested | Characterized | Sequenced | Sent |
|---|---|---|---|---|---|---|---|
| Final Constructs | (!) Favorite Part | ||||||
| Final Prey Cell (!) | J37015 | YES | YES | Pending | YES | YES | |
| | Cre/Lox Prey Control (!) | J37027 | YES | YES | YES | YES | YES |
| Test Constructs | |||||||
| Final Polycistronic Predator Cell Test Construct (!) | J37016 | YES | YES | YES | YES | YES | |
| | Predator Cell, pLux Transfer Function (two promoters) | J37020 | YES | YES | NO | NO | YES |
| Intermediate Parts | |||||||
| | AHL induced LuxR generator (for predator cell) | J37019 | YES | N/A | N/A | YES | YES |
| | RBS + LuxR | J37033 | YES | N/A | N/A | YES | YES |
| | Prey Cell Intermediate | J37034 | YES | N/A | N/A | YES | YES |
| | LuxR + GFP | J37032 | YES | N/A | N/A | YES | YES |
Our Open Documentation
- 300+ Pages of documentation on OWW, 100's of Photographs, Extensive Modelling, Lab Notebook and much more!
|
|
|
|
|---|
The Team and Acknowledgements
Undergrads
|
Advisors
|
Funding
|
|---|