ETH Zurich 2006
From 2006.igem.org
Christophe (Talk | contribs) (→Design of the Counter) |
Christophe (Talk | contribs) |
||
| Line 103: | Line 103: | ||
== [[Input-module]] == | == [[Input-module]] == | ||
| - | + | The input module splits the input into two opposite signals. It is best described through its system boundaries. One of the outputs should be high and the other low when S is high and vice versa when S is low: | |
| + | |||
| + | [[Image:inputPops.png]] | ||
| - | + | === Biological Implementation === | |
| - | [[Image: | + | To achieve such behaviour, we use the λ-system, with IPTG as inductor. It is relatively easy to handle/debug, and does not restrict the module from being extended to work with other types of inputs. More importantly, it is already available as a BioBrick (Registry package 7.05) in its unidirectional flavour (In nature, the λ-system is bidirectional, with Pr on one DNA strand and Prm on the other, overlapping). The following picture shows the gene circuit of the input module in details: |
| + | |||
| + | [[Image:Lambda-Sys in INPUT-moduleSmall.jpg|450px|Modified Lambda-system in INPUT-module: unidirectional, no OR3]] | ||
| + | |||
| + | cI is a dimer and regulates the activity of the two promoter regions, Pr and Prm, on the λ-system. Pr is constitutively active and is repressed when cI binds to the two operator regions it overlaps with (OR1, OR2). Conversly, Prm has low basal activity, and is activated by cI. Since the two promoters are regulated by the same protein-operator interactions, repression and activation is expected to be symmetrical (a necessary condition, see results from simulation below). For more details, please consult the page [[Input-module]]. | ||
== [[NOR-module|NOR Module]] == | == [[NOR-module|NOR Module]] == | ||
Revision as of 08:00, 21 October 2005
Short Overview. The project of the ETH Zurich team consists of the design and implementation in vivo of a gene circuit that can count up to 4. In essence, the counter uses two toggle switches, each storing 1 bit, to keep track of the 4 states. The design of the counter is highly modular, with the hope that it can be included as a unit in larger circuits, and also combined with further counter instances to keep track of a much larger number of states, up to (2^(n+1)) with n units. To facilitate further developments and integration to other projects, the counter is available in form of BioBricks. Among many exciting applications, the availability of a counter enables the execution of sequential instructions, and paves the way for the execution of artifical programs inside living cells.
Contents |
News
- 2005.10.18 The parts for the actual INPUT-module are ready, thanks to the hard work of Giorgia, Hervé, and Martje (not all test/debbuging parts though)
- 2005.10.07 Message from Blue Heron: Sequences for NOR are synthesized and will be verified and assembled next week.
- 2005.09.23 Sequences for NOR module ordered from Blue Heron
Organisation
People
Students
| Simon Barkow | Christophe Dessimoz | Zlatko Franjcic |
| Dominic Frutiger | Robin Künzler | Urs A. Müller |
| Jonas Nart | Kristian Nolde | Alexander Roth |
| Tamara Ulrich | Giorgia Valsesia | Herve Vanderschuren |
Supervisors
| Jörg Stelling | Sven Panke | Eckart Zitzler |
Advisors
| Uwe Sauer | Martin Fussenegger | Andreas Hierlemann |
| Kay-Uwe Kirstein | Ruedi Aebersold |
Timeline & Tasks
Groups
Modeling
- Members: Tamara, Kristian, Zlatko
This group is focused on mathematical modeling, illustration, simulations and overall design. There are no permanent members for this group. Anyone can join and leave the group anytime he/she wants. A group coordinator will be assigned at the first meeting.
- Meetings:
empty
Input Module
- Members: Christophe, Dominic (coordinator), Giorgia, Herve, Zlatko
- Meetings:
log 2005-08-17: Wednesday, 13:00 @ polyterrasse: Discussion of module, next steps, task distrib. log 2005-08-22: Monday, 15:00 @ polyterrasse: Discussion of biol. solutions. log 2005-09-12: Thursday, 17:30 teammeeting. ... multiple meetings, no log ... 2005-10-10: Monday, 10:00 @ intro to FACS system (Sven, Giorgia, Hervé, Dominic). 2005-10-11: Tuesday, 15:00 @ polyterrasse: planning test session (Giorgia, Dominic) log 2005-10-13: Thursday, 17:30 @ test session at Leica AOBS at LMC (Giorgia, Dominic) 2005-10-19: Wednesday, 14:00 @ planning of tests, giorgia, hervé, dominic)
NOR Module
- Members: Alex (coordinator), Jonas, Robin, Simon, Tamara, Urs
- Meetings:
log 2005.08.17, Wednesday, 15:30 2005.08.18, Thursday, 13:15 log 2005.08.22, Monday, 13:45 2005.08.25, Thursday 15:30 2005.08.29, Monday 14:00 2005.08.30, Tuesday 16:00 2005.09.06, Tuesday 08:00
Documentation: 'todo' list
Introduction
The past few years have seen the emergence of the field of synthetic biology, in which functional units are designed and built into living cells to generate a particular behaviour, and ultimately to better understand Life's mechanisms. Previous efforts include the creation of gene circuits that generate oscillating behaviour (Elowitz00), toggle switch functionality (Atkinson03), artificial cell-cell communication (Bulter04) or pattern-forming behaviour (Basu2005). The present document describes the design and realization of a gene circuit that counts to 4.
Modeling
Design of the Counter
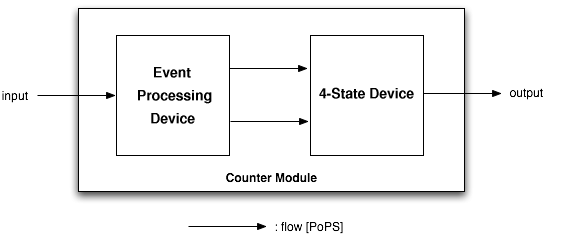
The counter is a genetic circuit that has 1 input and 4 outputs. It uses the input signal to switch from one of the four output to the next. When the input signal is high, either output 1 or 3 is active, when it is low, output 2 or 4 is active. Thus, output 1 and 3 alternatively keep track of high input signal, while output 2 and 4 alternatively keep track of low input signals.
As depicted above, the counter is made of two parts, serially linked:
- the "Input" module, which splits the input into two opposite signals.
- the "NOR" module, which uses these two signals to sequencially switch through the outputs 1, 2, 3 and 4.
Note that all interfaces have flows described in Polymerase Per Second (PoPS), is explained in details on the [http://partsregistry.org/cgi/htdocs/AbstractionHierarchy/index.cgi abstraction hierarchy] of the MIT Registry of Parts. For instance, the input can be of any nature as long as an adequate promoter is available (e.g. heat-shock using a sigma32 promoter, IPTG using a LacI promoter, AHL using quorum sensing promoters...)
Input-module
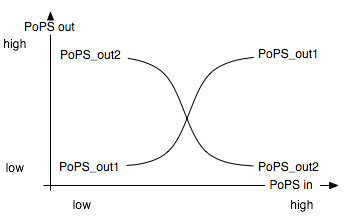
The input module splits the input into two opposite signals. It is best described through its system boundaries. One of the outputs should be high and the other low when S is high and vice versa when S is low:
Biological Implementation
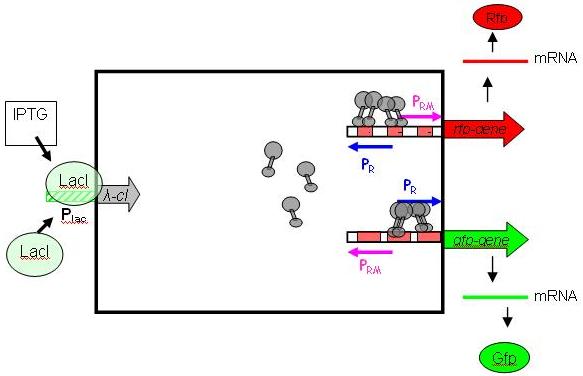
To achieve such behaviour, we use the λ-system, with IPTG as inductor. It is relatively easy to handle/debug, and does not restrict the module from being extended to work with other types of inputs. More importantly, it is already available as a BioBrick (Registry package 7.05) in its unidirectional flavour (In nature, the λ-system is bidirectional, with Pr on one DNA strand and Prm on the other, overlapping). The following picture shows the gene circuit of the input module in details:
cI is a dimer and regulates the activity of the two promoter regions, Pr and Prm, on the λ-system. Pr is constitutively active and is repressed when cI binds to the two operator regions it overlaps with (OR1, OR2). Conversly, Prm has low basal activity, and is activated by cI. Since the two promoters are regulated by the same protein-operator interactions, repression and activation is expected to be symmetrical (a necessary condition, see results from simulation below). For more details, please consult the page Input-module.
NOR Module
A NOR-module works as follows: There are two inputs and one output. The output is high if and only if the two inputs are low. In other words: The output is low whenever either one (or both) of the inputs are high.
Results
Discussion
Appendix
References
Modeling and illustration tools,
bulter04, atkinson03, bates05, keiler01, suetsugu03, sudesh00, römling02, ross91, sutherland01, Lai04, zogaj01, miller01, basu05, goryachev05, you04, Isalan01, Beerli02, Mani05, Beerli00, Beerli98, Dreier01, Dreier05, Klug05, Yang95, Segal99, Segal03,
Glossary
Previous Ideas
This is the brainstorming and previous ideas section. In this section you will find other projects that had been pursued, as well as random ideas without too much consideration of feasibility, etc.