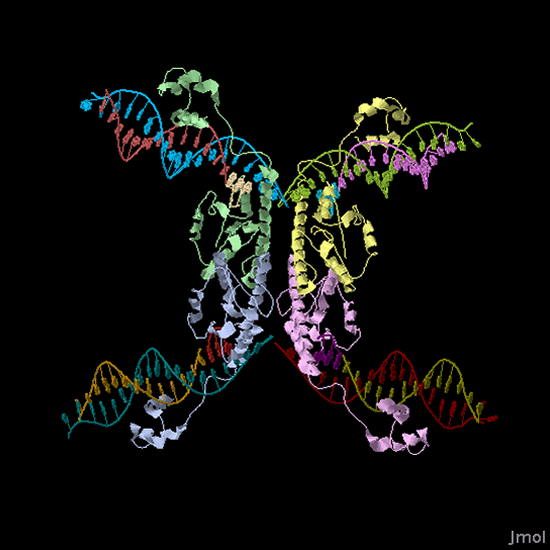Davidson 2006
From 2006.igem.org
(→'''Project Description''') |
|||
| (369 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | <center>[[Image:logo.gif]]</center> | + | {| cellpadding="10" width="900px" |
| + | |- valign="top" | ||
| + | | <center>[[Image:logo.gif]]<br>http://www.davidson.edu<br><br><big>'''[[#overview|Project Overview]]'''<br><br>'''[http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Davidson Davidson Parts]'''<br><br>'''[[#members|Team Members]]'''<br><br>'''[[#tools|Tools and Resources]]'''<br><br>Check out our [http://www.bio.davidson.edu/courses/synthetic/photos/FlapJack_HotCakes.html Official Team Photo]<br><br> | ||
| + | <font color = red>NEW</font> [[Davidson's Awards]] and [[Photo Gallery]] from iGEM Jamboree 2006</big></center> | ||
| + | | [[Image:igem_team_web.jpg|thumb|450px|Left to Right: Lance, Erin, Samantha, Sabriya, Karmella, Laurie, Malcolm]] | ||
| + | |} | ||
| + | |||
| + | ---- | ||
| + | {| width="900px" | ||
| + | |- valign="top" | ||
| + | |<font size=4><div id="overview">Solving the Pancake Problem with an ''E. coli'' Computer</div></font> | ||
| + | [http://2006.igem.org/Image:IGEM2006T_Davidson.ppt PowerPoint Presentation] and [http://2006.igem.org/Image:IGEM2006_Davidson_poster.ppt Poster] | ||
| + | |- | ||
| + | |Our goal is to genetically engineer a biological system that can compute solutions to a puzzle called the burnt pancake problem. The '''E.HOP computer''' is a proof of concept for computing ''in vivo,'' with implications for future data storage devices and transgenic systems. Our work was done in collaboration with the [http://2006.igem.org/wiki/index.php/Missouri_Western_State_University_2006 Missouri Western iGEM Team] and an undergraduate research fellow from [http://www.hamptonu.edu/ Hampton University]. | ||
| + | |[[image:EHOP.gif|250px]] | ||
| + | |} | ||
| - | == ''' | + | ====The Burnt Pancake Problem==== |
| + | {| width="900px" | ||
| + | |- valign="top" | ||
| + | |The [http://en.wikipedia.org/wiki/Pancake_sorting pancake problem] is a puzzle in which a scrambled series of units (or stack of pancakes) must be shuffled into the correct order and orientation. You can try solving [http://www.cut-the-knot.org/SimpleGames/Flipper.shtml a simple version of the pancake problem] yourself.<br><br>In the burnt pancake problem, each pancake is given an orientation by burning one side. To solve the problem, every unit, or pancake, must be placed in the proper order (largest on bottom, smallest on top) and in the proper orientation (burnt side down, golden side up). The pancakes are flipped with two spatulas: one to lift pancakes off the top of the stack, the other to flip part (or all) of the remaining stack of pancakes. The pancakes lifted by the first spatula are returned to the top of the stack without being flipped. Figure 1 illustrates how the puzzle is solved. | ||
| + | |- | ||
| + | |[[Image:solving4cakes.jpg|thumb|600px|center|'''Figure 1.''' A scrambled stack of four burnt pancakes, sorted into correct order and orientation in 3 flips. Proper orientation is indicated by solid color, improper orientation by hatched color. ]] | ||
| + | |} | ||
| - | + | ==== Approach ==== | |
| + | {| width="900px" | ||
| + | |- valign="top" | ||
| + | | Trial and error is one approach to solving the burnt pancake problem, but how could one compute the quickest solution? Our idea is to let E. coli do the work, using each cell as a tiny processor in a massively parallel machine. A mathematical model of the flipping process helps us design the system and interpret the output of our EHOP computer. | ||
| + | |- valign="top" | ||
| + | |[[Image:Jmol_Hin_tetrad_DNA.gif|thumb|250px|left|'''Figure 2.''' 3-D structure of a Hin protein complex bound to DNA. View the [http://www.rcsb.org/pdb/explore/explore.do?structureId=1ZR4 interactive 3-D Jmol image].]]'''Biological System:''' The biological representation of a pancake is a functional unit of DNA such as a promoter or coding region. To flip these units of DNA, we have reconstituted the Hin/ Hix invertase system (Figure 2) from ''Salmonella typhimurium'' as a BioBrick compatible system in ''E. coli''. '''Hin invertase''' (<partinfo>J31001</partinfo>) was cloned from ''S. typhimurium'', Ames strain TA100 and tagged with LVA. We built the recombinational enhancer (RE) and Hin invertase recognition sequence HixC using the publicly available genomic sequence of ''S. typhimurium'' and [http://gcat.davidson.edu/IGEM06/oligo.html a dsDNA assembly program we created] for gene synthesis from overlapping oligos. | ||
| - | + | Every segment of DNA flanked by a pair of HixC sites is a "pancake" capable of being inverted. Hin invertase recognizes pairs of HixC sites and inverts the DNA fragment in between the two HixC sites with the help of the Fis protein bound to the RE. In our system, selectable phenotypes (including antibiotic resistance and RFP expression), depend upon the proper arrangement of a series of HixC-flanked DNA segments in a plasmid. This allows us to select for cells that have successfully solved the puzzle. An example of a sorted stack of two pancakes is shown in Figure 3. There are 7 other configurations of 2 pancakes. If we increase the number of pancakes to 4, there are 384 configurations. In general, there are 2<sup>n</sup> n! configurations of ''n'' pancakes, a combinatorial explosion of possibilities. How many should we build and test? Which ones are most informative? We need a mathematical model to help answer this question. | |
| - | + | |- valign="top" | |
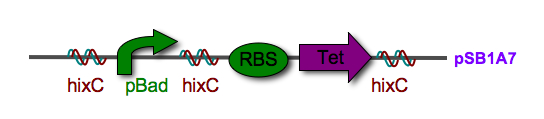
| + | |[[Image:biocake.jpg|thumb|400px|right|'''Figure 3.''' A sorted stack of two biological "pancakes"]] | ||
| - | + | '''Mathematical Model:''' Our mathematical representation of a stack of pancakes is a '''signed permutation''', in which each numerical value represents the pancake size and the sign of the number represents the orientation. For example, (1, 2, 3, 4) is a sorted stack of four pancakes, all in the proper order and orientation, and (-2, 4, -1, 3) is the scrambled stack of four pancakes shown on the far left of Figure 1. Note that pancakes 1 and 2 are negative, indicating that these two pancakes are in the wrong orientation (burnt side up). | |
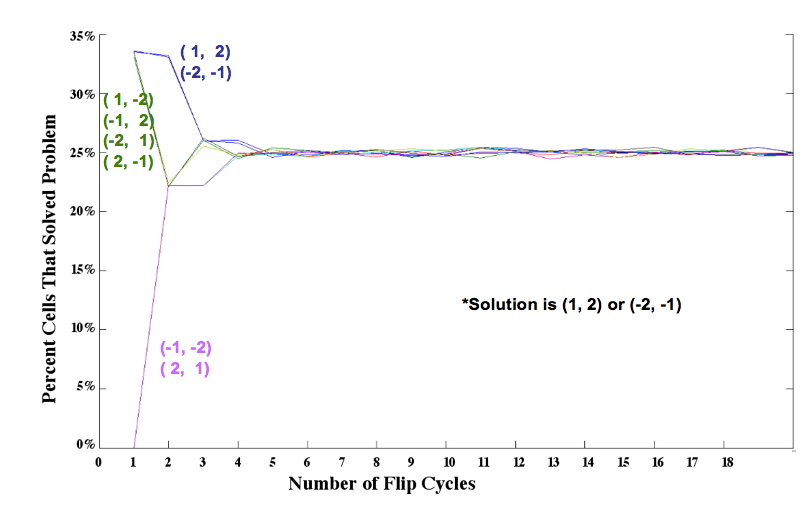
| - | + | [[Image:two_pancake_families.jpg|thumb|400px|right|'''Figure 4.''' Simulation results for two pancakes]] We simulated the flipping process by assuming that each segment of DNA flanked by Hix sites is equally likely to be flipped by Hin invertase. In an ''n''-pancake stack, there are <sub>n+1</sub>C<sub>2</sub> ways to choose a segment: two spatula locations must be chosen from among the ''n'' + 1 potential locations. For example, there are <sub>5</sub>C<sub>2</sub> = 5! 3! / 2! = 10 different ways to perform a single flip in a stack of 4 pancakes. The random flipping process continues until the stack is sorted in the correct order and orientation, or until we artificially stop the random flipping process. Figure 4 shows the percent of plasmids that we expect to be sorted after 1 flip, 2 flips, and so on, up to 20 flips, for each of the eight initial configurations. | |
| - | + | From these simulations, and similar results for stacks of 3 and 4 pancakes, we discovered that the probability of a plasmid being sorted after ''k'' flips was determined by how many ways the solution could be obtained. Several initial configurations had essentially the same graph in our simulations because they had the same number of ways of being sorted. Only '''one''' representative from each of these "families" of initial configurations needs to be built and tested. | |
| + | |} | ||
| + | {| width="900px" cellpadding="5" border="1" border-style="solid" border-color="blue" | ||
| + | |- | ||
| + | | align="center" colspan="4"| <big>'''Methods and Results'''</big><br>The following is an outline of what the Davidson team will present at iGEM 2006 | ||
| + | |- valign="top" | ||
| + | | width="300px" | <big>'''Basic parts'''</big><br><br> | ||
| + | Parts used in this project were designed by two iGEM Teams | ||
| + | *[http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Davidson Davidson] | ||
| + | *[http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Missouri Missouri Western]<br><br> | ||
| + | <big>'''Modeling'''</big><br><br> | ||
| + | *[[Modeling pancake flipping]]: deducing kinetics and size biases | ||
| + | *[[Choosing constructs]]: deciding which unsorted pancake stacks to start with | ||
| + | *[[The effect of plasmid copy number]] | ||
| + | | width="300px" | <big>'''Building the Biological System'''</big><br><br> | ||
| + | * [[DNA Pancake|Defining a DNA pancake]] - what parts are flippable? | ||
| + | * [[Read Through Transcription Problem|Read-through from the carrier vector backbone]] leads to uncontrolled Tet expression and uncontrolled flipping | ||
| + | * [[pSB1A7| New vector insulates parts from read-through]], but is not compatible with parts carrying double terminator B0015 | ||
| + | * [[pSB1A7 Cloning Solution| Cloning solution]] - design pancake constructs without double terminator B0015 | ||
| + | * [[Biological Equivalence Problem|Dealing with biological equivalence]] - distinguishing 1,2 from -2,-1 using an RFP reporter | ||
| + | * [[Two Pancake Stacks|Two-pancake stacks]] - final design | ||
| + | | width="300px" | <big>'''Conclusions'''</big><br><br> | ||
| + | *'''[[Consequences of devices]]''' | ||
| + | ** Practical: proof-of-concept for bacterial computers, data storage | ||
| + | ** Basic research: transgene rearrangement in vivo, insights into evolution that has occurred via DNA rearrangements | ||
| + | *'''[[Next steps]]''': can solve problem but need control over kinetics | ||
| + | *'''[[Lessons learned]]''': | ||
| + | **Troubleshooting, communication, teamwork, publicity | ||
| + | **Math and Biology meshed really well and even uncovered a new proof | ||
| + | **Multiple campuses can increase capacity through communication and cooperation | ||
| + | **Size of school is not a limiting factor | ||
| + | **We had a blast and learned heaps!!! | ||
| + | |} | ||
| + | ---- | ||
| + | <font size=4><div id="members">TEAM MEMBERS</div></font> | ||
| - | + | '''Students''' | |
| + | * [mailto:fizzle6821@yahoo.com| Sabriya Rosemond] is a junior biology major at Hampton University. | ||
| + | * [mailto:erzwack@davidson.edu| Erin Zwack] is a junior biology major at Davidson College. | ||
| + | * [mailto:laharden@davidson.edu| Lance Harden] is a sophomore math major at Davidson College. | ||
| + | * [mailto:sasimpson@davidson.edu| Samantha Simpson] is a sophomore at Davidson College who might design a major in genomics. | ||
| - | + | '''Faculty''' | |
| + | * [mailto:macampbell@davidson.edu| A. Malcolm Campbell] [http://www.bio.davidson.edu/campbell Department of Biology] | ||
| + | * [mailto:laheyer@davidson.edu| Laurie J. Heyer] [http://www.davidson.edu/math/heyer/ Department of Mathematics] | ||
| + | * [mailto:kahaynes@davidson.edu| Karmella Haynes] [http://www.bio.davidson.edu/ Department of Biology - teaching postdoc and visiting assistant professor] | ||
| - | + | ---- | |
| - | + | <font size=4><div id="tools">Tools and Resources</div></font> | |
| - | + | {| width="840px" cellpadding="5" border="0" | |
| - | + | |- valign="top" | |
| - | + | | width="210px" | <big>'''iGEM 2006 Jamboree'''</big><br><br> | |
| - | + | *[[Presentation Outline]] | |
| - | + | *[http://2006.igem.org/Image:IGEM2006T_Davidson.ppt PowerPoint Presentation] | |
| - | + | *[http://2006.igem.org/Image:IGEM2006_Davidson_poster.ppt Poster] - ppt file | |
| - | + | | width="210px" | <big>'''White Board'''</big><br><br> | |
| - | + | * [[General Questions]] | |
| - | < | + | * [[Math Questions]] |
| - | + | * [[Biology Questions]] | |
| - | + | | width="210px" | <big>'''Assembly Plans'''</big><br><br> | |
| - | <br> | + | *[[Assembly powerpoint]] |
| - | <br> | + | *[[Assembly plan]] |
| - | + | *[[Western Assembly Plan]] | |
| - | + | *[[Davidson Assembly Plan]] | |
| - | + | *[[pLac Tet Pancake Plan]] (new) | |
| - | + | | width="210px" | <big>'''Progress'''</big><br><br> | |
| - | + | *[[Plasmids and Bacteria]] | |
| - | < | + | *[[Hix and RE]] |
| - | + | *[[Hin and Antibiotic Pancakes]] | |
| - | + | *[[Ligations]] | |
| - | + | |- valign="top" | |
| - | + | | width="210px" | <big>'''Wet Lab Tools'''</big><br><br> | |
| - | + | *[http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Davidson Davidson Parts] | |
| - | + | *[http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Missouri Missouri Western Parts] | |
| - | + | *Summer 2006 [[Pancake Parts]] | |
| - | + | *[[The What's and How's| Davidson Protocols]] | |
| - | + | *[http://gcat.davidson.edu/IGEM06/oligo.html Oligo Cuts Optimization]: by Lance Harden | |
| - | + | *[[Papers of Interest]] | |
| - | + | *[[Freezer Stocks]] | |
| - | + | | width="210px" | <big>'''Simulators'''</big><br><br> | |
| - | ''' | + | *<font color='red'>T</font color><font color='blue'>h</font color><font color='orange'>e</font color><font color='silver'> L</font color><font color='sky blue'>a</font color><font color='magenta'>n</font color><font color= 'orange'>c</font color><font color='green'>e</font color><font color=cyan>l</font color><font color='red'>a</font color><font color='brown'>t</font color><font color='blue'>o</font color><font color='violet'>r</font color> Pancake Flipping Simulator(MATLAB Code) |
| - | + | * Other Flipping Simulators | |
| - | + | **[http://gcat.davidson.edu/IGEM06/randompancakeflipper.m User-Friendly Version] | |
| - | + | **[http://gcat.davidson.edu/IGEM06/powerflipper.m Power-User Version] | |
| - | + | **[http://gcat.davidson.edu/IGEM06/powerflipperplus.m Power-User Update] | |
| - | * | + | **[http://gcat.davidson.edu/IGEM06/findmin.m Findmin] |
| - | * | + | **[http://gcat.davidson.edu/IGEM06/bigtrial.m Bigtrial] |
| - | * | + | | width="210px" | <big>'''Graphing Tools'''</big><br><br> |
| - | * | + | *[http://gcat.davidson.edu/IGEM06/pancakeplotter.m Pancakeplotter] |
| - | * | + | *[http://gcat.davidson.edu/IGEM06/permplotter.m Permplotter] |
| - | + | *[http://gcat.davidson.edu/IGEM06/adjmat.m Adjmat - creates adjacency matrices] | |
| - | ''' | + | *[http://gcat.davidson.edu/IGEM06/bioadjmat.m Bioadjmat - Adjmat with biological equivalence] |
| - | + | *[http://gcat.davidson.edu/IGEM06/make_radial.m Make_radial - creates a radially-embedded graph] | |
| - | + | *[http://gcat.davidson.edu/IGEM06/find_diam.m Find_diam - finds the diameter of a pancake graph] | |
| - | + | | width="210px" | <big>'''Miscellaneous Computation Tools'''</big><br><br> | |
| - | + | *[http://gcat.davidson.edu/IGEM06/signedperms.m Signedperms - lists all signed permutations for a stack of k pancakes] | |
| - | + | *[http://gcat.davidson.edu/IGEM06/bioperms.m Bioperms - Signedperms with biological equivalence] | |
| - | + | |} | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | ''' | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Latest revision as of 19:12, 19 February 2007
 http://www.davidson.edu Project Overview [http://partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006partsregistry.org/cgi/partsdb/pgroup.cgi?pgroup=iGEM2006&group=Davidson Davidson Parts] Team Members Tools and Resources Check out our [http://www.bio.davidson.edu/courses/synthetic/photos/FlapJack_HotCakes.html Official Team Photo] NEW Davidson's Awards and Photo Gallery from iGEM Jamboree 2006 |
The Burnt Pancake Problem
| The [http://en.wikipedia.org/wiki/Pancake_sorting pancake problem] is a puzzle in which a scrambled series of units (or stack of pancakes) must be shuffled into the correct order and orientation. You can try solving [http://www.cut-the-knot.org/SimpleGames/Flipper.shtml a simple version of the pancake problem] yourself. In the burnt pancake problem, each pancake is given an orientation by burning one side. To solve the problem, every unit, or pancake, must be placed in the proper order (largest on bottom, smallest on top) and in the proper orientation (burnt side down, golden side up). The pancakes are flipped with two spatulas: one to lift pancakes off the top of the stack, the other to flip part (or all) of the remaining stack of pancakes. The pancakes lifted by the first spatula are returned to the top of the stack without being flipped. Figure 1 illustrates how the puzzle is solved. |
Approach
| Trial and error is one approach to solving the burnt pancake problem, but how could one compute the quickest solution? Our idea is to let E. coli do the work, using each cell as a tiny processor in a massively parallel machine. A mathematical model of the flipping process helps us design the system and interpret the output of our EHOP computer. |
Biological System: The biological representation of a pancake is a functional unit of DNA such as a promoter or coding region. To flip these units of DNA, we have reconstituted the Hin/ Hix invertase system (Figure 2) from Salmonella typhimurium as a BioBrick compatible system in E. coli. Hin invertase () was cloned from S. typhimurium, Ames strain TA100 and tagged with LVA. We built the recombinational enhancer (RE) and Hin invertase recognition sequence HixC using the publicly available genomic sequence of S. typhimurium and [http://gcat.davidson.edu/IGEM06/oligo.html a dsDNA assembly program we created] for gene synthesis from overlapping oligos.
Every segment of DNA flanked by a pair of HixC sites is a "pancake" capable of being inverted. Hin invertase recognizes pairs of HixC sites and inverts the DNA fragment in between the two HixC sites with the help of the Fis protein bound to the RE. In our system, selectable phenotypes (including antibiotic resistance and RFP expression), depend upon the proper arrangement of a series of HixC-flanked DNA segments in a plasmid. This allows us to select for cells that have successfully solved the puzzle. An example of a sorted stack of two pancakes is shown in Figure 3. There are 7 other configurations of 2 pancakes. If we increase the number of pancakes to 4, there are 384 configurations. In general, there are 2n n! configurations of n pancakes, a combinatorial explosion of possibilities. How many should we build and test? Which ones are most informative? We need a mathematical model to help answer this question. |
|
Mathematical Model: Our mathematical representation of a stack of pancakes is a signed permutation, in which each numerical value represents the pancake size and the sign of the number represents the orientation. For example, (1, 2, 3, 4) is a sorted stack of four pancakes, all in the proper order and orientation, and (-2, 4, -1, 3) is the scrambled stack of four pancakes shown on the far left of Figure 1. Note that pancakes 1 and 2 are negative, indicating that these two pancakes are in the wrong orientation (burnt side up). We simulated the flipping process by assuming that each segment of DNA flanked by Hix sites is equally likely to be flipped by Hin invertase. In an n-pancake stack, there are n+1C2 ways to choose a segment: two spatula locations must be chosen from among the n + 1 potential locations. For example, there are 5C2 = 5! 3! / 2! = 10 different ways to perform a single flip in a stack of 4 pancakes. The random flipping process continues until the stack is sorted in the correct order and orientation, or until we artificially stop the random flipping process. Figure 4 shows the percent of plasmids that we expect to be sorted after 1 flip, 2 flips, and so on, up to 20 flips, for each of the eight initial configurations.From these simulations, and similar results for stacks of 3 and 4 pancakes, we discovered that the probability of a plasmid being sorted after k flips was determined by how many ways the solution could be obtained. Several initial configurations had essentially the same graph in our simulations because they had the same number of ways of being sorted. Only one representative from each of these "families" of initial configurations needs to be built and tested. |
| Methods and Results The following is an outline of what the Davidson team will present at iGEM 2006 | |||
| Basic parts Parts used in this project were designed by two iGEM Teams
Modeling
| Building the Biological System
| Conclusions
| |
Students
- Sabriya Rosemond is a junior biology major at Hampton University.
- Erin Zwack is a junior biology major at Davidson College.
- Lance Harden is a sophomore math major at Davidson College.
- Samantha Simpson is a sophomore at Davidson College who might design a major in genomics.
Faculty
- A. Malcolm Campbell [http://www.bio.davidson.edu/campbell Department of Biology]
- Laurie J. Heyer [http://www.davidson.edu/math/heyer/ Department of Mathematics]
- Karmella Haynes [http://www.bio.davidson.edu/ Department of Biology - teaching postdoc and visiting assistant professor]
iGEM 2006 Jamboree
| White Board | Assembly Plans | Progress |
Wet Lab Tools
| Simulators
| Graphing Tools
| Miscellaneous Computation Tools
|