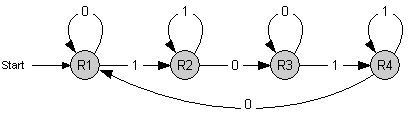ETH Zurich 2006
From 2006.igem.org
(→Biological Implementation) |
m (→References: removed two links) |
||
| Line 209: | Line 209: | ||
[[Isalan01]], | [[Isalan01]], | ||
[[keiler01]], | [[keiler01]], | ||
| - | |||
[[Klug05]] (Minireview), | [[Klug05]] (Minireview), | ||
[[Lai04]], | [[Lai04]], | ||
[[Mani05]], | [[Mani05]], | ||
[[miller01]], | [[miller01]], | ||
| - | |||
[[Newman03]] (Leucine Zippers), | [[Newman03]] (Leucine Zippers), | ||
[[Park et al.]] 2005 (Activation of transcription), | [[Park et al.]] 2005 (Activation of transcription), | ||
Revision as of 11:50, 28 October 2005
Contents |
News
- 2005.10.25 Now we know that Blue Heron will not be able to deliver in time.
- 2005.10.24 We had a very intensive meeting on the overall concept and the documentation of the project. This page will change a lot in the coming days.
- 2005.10.23 One month after ordering, the sequences are still not ready. That means it is unlikely for us to complete the 4-State Device before the jamboree unless we find some alternative solutions (and have quite a bit of luck). See message from blue herons web page:
IMPORTANT NOTE: Despite Blue Heron Bio's capacity expansion, record order volume has created larger than anticipated queuing, which has increased delivery times for some orders. We are reviewing Estimated Ship Dates regularly to provide you with our best estimate of when your order will ship. We are working hard to deliver your orders as quickly as possible, and we are implementing new capacity management systems to avoid this problem in the future. If you have any questions or concerns please call our Customer Service.
- 2005.10.18 The parts for the actual Event Processing Device are ready, thanks to the hard work of Giorgia, Hervé, and Martje (not all test/debugging parts though)
- 2005.10.07 Message from Blue Heron: Sequences for NOR are synthesized and will be verified and assembled next week.
- 2005.09.23 Sequences for 4-State Device ordered from Blue Heron
Organisation
People
Students
| Simon Barkow | Christophe Dessimoz | Zlatko Franjcic |
| Dominic Frutiger | Robin Künzler | Urs A. Müller |
| Jonas Nart | Kristian Nolde | Alexander Roth |
| Tamara Ulrich | Giorgia Valsesia | Herve Vanderschuren |
Supervisors
| Jörg Stelling | Sven Panke | Eckart Zitzler |
Advisors
| Uwe Sauer | Martin Fussenegger | Andreas Hierlemann |
| Kay-Uwe Kirstein | Ruedi Aebersold |
Timeline
Tasks
=Abstract= The project of the ETH Zurich team consists of the design and implementation [http://en.wikipedia.org/wiki/In_vivo in vivo] of a gene circuit that can count to 2. In essence, the counter uses two toggle switches, each storing 1 bit, to keep track of the 4 internal states. The design of the counter is highly modular, with the hope that it can be included as a unit in larger circuits, and also combined with further counter instances to keep track of a much larger number of states, up to 2^n with n units. To facilitate further developments and integration to other projects, the counter is available in form of [http://parts2.mit.edu| BioBricks]. Among many exciting applications, the availability of a counter enables the execution of sequential instructions, and paves the way for the execution of artificial programs inside living cells.
Introduction
The past few years have seen the emergence of the field of Synthetic Biology, in which functional units are designed and built into living cells to generate a particular behaviour, and ultimately to better understand Life's mechanisms. Previous efforts include the creation of gene circuits that generate oscillating behaviour (Elowitz00), toggle switch functionality (Atkinson03), artificial cell-cell communication (Bulter04) or pattern-forming behaviour (Basu2005). The present document describes the design and realization of a gene circuit as a basic device that counts external events to 2.
Concept
The counter is a finite state machine implemented as a genetic circuit. It has 4 internal states s1 to s4. The transition between these states is induced by an external stimulus with values 0 and 1 - denoting whether it is absent or present, respectively. Repeated stimulus will lead to successive transitions and finally to repeated cycling through those 4 states.
Each time the state s3 is reached, an output signal is generated. This leads to a counting behavior where every second occurence of 1 (high signal) is indicated by the output signal.
Further information on the state machine
The coupling of further instances of this state machine allows to count to higher numbers, up to 2^n with n units.
System Implementation
System Architecture
System-level Diagram
Device-level Diagram
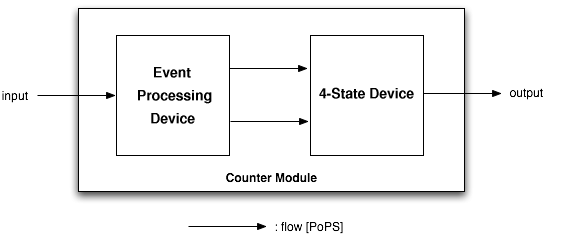
As depicted above, the counter is made of two parts, serially linked:
- The Event Processing Device, which splits the input into two outputs: one feed through and one inverted signal - which serve in turn as inputs for the 4-State Device.
- The 4-State Device module, which uses these two signals to sequentially switch through the states S1, S2, S3 and S4.
Note that all interfaces have flows described in Polymerase Per Second (PoPS), is explained in details on the [http://partsregistry.org/cgi/htdocs/AbstractionHierarchy/index.cgi abstraction hierarchy] of the MIT Registry of Parts. For instance, the input can be of any nature as long as an adequate promoter is available (e.g. heat-shock using a sigma32 promoter, IPTG using a LacI promoter, AHL using quorum sensing promoters...)
Parts-level Diagram
... parts with the typical illustration as found registry.
Concentration Dynamics Diagram
Devices
Event Processing Device
Basic Functionality
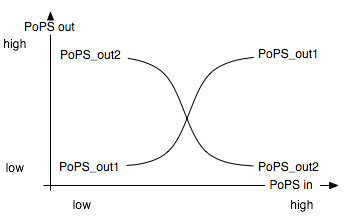
The Event Processing Device splits the input induced by an external event into two signals: one is basically fed through while the other is inverted. It is best described through its system boundaries. One of the outputs should be high and the other low when S is high and vice versa when S is low [Fig EPD.1]. Both output signals then serve as input signals for the 4-State Device and it is thus necessary that they have the same delay.
Biological Implementation
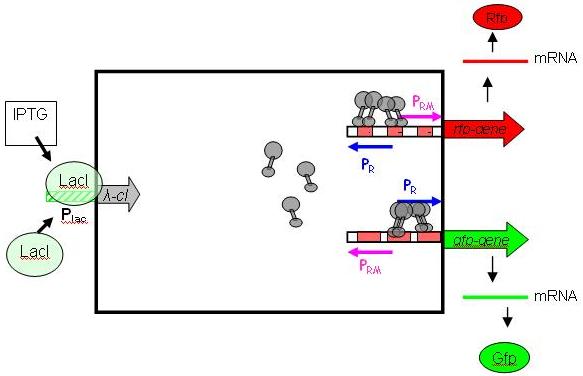
To achieve such behaviour, we use the [http://en.wikipedia.org/wiki/Lambda_phage λ phage system], with IPTG as inductor. It is relatively easy to handle/debug, and does not restrict the module from being extended to work with other types of inputs. More importantly, it is already available as a BioBrick (Registry package 7.05) in its unidirectional flavour (In nature, the λ phage system is bidirectional, with Pr on one DNA strand and Prm on the other, overlapping). The following picture shows the gene circuit of the Event Processing Device in detail [Fig EPD.2].
cI is a dimer and regulates the activity of the two promoter regions, Pr and Prm, on the λ phage system. Pr is constitutively active and is repressed when cI binds to the two operator regions it overlaps with (OR1, OR2). Conversely, Prm has low basal activity, and is activated by cI. Since the two promoters are regulated by the same protein-operator interactions, repression and activation is expected to be symmetrical (a necessary condition, see results from simulation below).
Detailed Documentation
For more details, please consult the main page for the Event Processing Device.
4-State Device
Basic Functionality
The 4-State Device uses two inputs to sequentially switch through its four states. Note that this behaviour can be observed abstractly in the concept section.
To achieve such behaviour in our system, we use four interconnected gates. In the following, we call them NOR gates. In essence, a NOR gate is a component that has two inputs and one output, where the output is only high if none of the input is high. Concretely, a NOR gate can be implemented through a promoter with high basal activity that is repressed by two effectors.
Our NOR gates have the two repression sites as described above and, additionally, one repression or one activation site. After each of the four NOR gates, there is a coding region for a repressor protein. These four different proteins are called R1 to R4 in the following.
(picture of one NOR gate, with a legend)
The following diagram shows how the four NOR gates are interconnected:
R3 R2 R4 R1 R4 R3 R1 R2
| | | | | | | |
/|____ | | | | ____|\ /|____ | | | | ____|\
___/ R1 |_=_=__ ___=_=_| R3 \___ ___/ R2 |_=_=___ __=_=_| R4 \___
\ ____| \ / |____ / \ ____| \ / |____ /
\| ^ ^ |/ \| ^ ^ |/
| | | |
| | | |
input 1 [PoPS] input 2 [PoPS]
By design of the Event Processing Device, input 1 and input 2 have opposite activity, meaning that either R1/R3 or R2/R4 is active. Furthermore, since R1/R3 (respectively R2/R4) are repressing each other, only one of the two is active. Therefore, in a stable situation, only 1 of the 4 repressor proteins is expressed.
Let us assume that R1 is being expressed. Input 2 must then be low, and therefore input 1 high. This situation is stable and remains until there is a change in the inputs. Now, if input 1 decreases, and input 2 increases, the expression of R1 will come to a halt. Since input 1 is now low, either R2 or R4 will be expressed. At this stage, R1 is still present in relatively high concentration and by repressing R4, it tips the balance in favor of R2, leading to a new stable state in which only R2 is expressed.
Note that electrical engineers call such a device a "J-K flip flop". It can also be seen as a combination of two toggle switches (Atkinson03), each being able to store one bit.
Biological Implementation
We use Zink-fingers proteins (ZFP) as repressors. This class of proteins binds to specific base pairs on the DNA. Many protein-DNA interaction for ZF domains and triplet of base pairs have been described, therefore making it possible to construct artificial transcription factors by combining ZF domains in a modular fashion. The idea is to use a ZFP as a repressor by putting a binding site for a ZFP upstream of the coding region and thereby preventing RNA polymerase to transcribe the gene.
Detailed Documentation
More details can be found in the dedicated 4-State Device page.
Mathematical Modeling
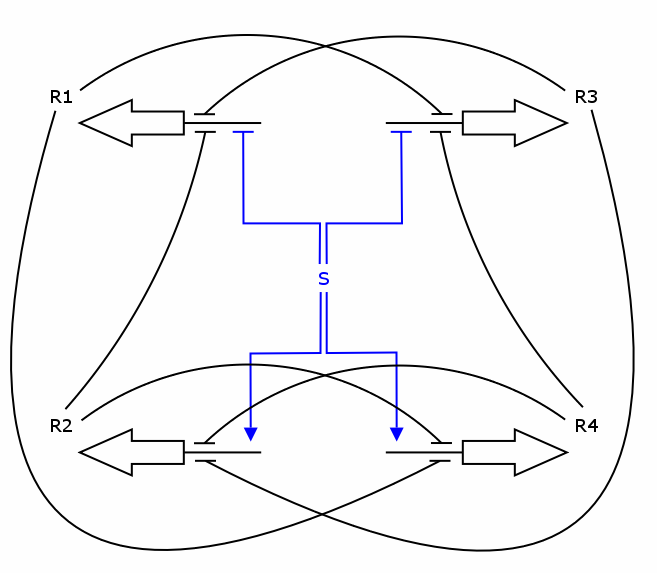
The simulation was performed through a deterministic model using ordinary differential equations (ODEs), as this approach is commonly used in modeling gene networks. Recall the counter architecture in [Figure Simulation 1].
R1 to R4 are the zincfingers that are expressed. S is the input of the counter. Note that the work of the Event Processing Device is symbolized by the dual effect of S, once as an activator and once as a repressor. Zincfingers R1 and R3 have Pr as a promoter, R2 and R4 have Prm as a promoter. See Event_Processing_Device for more details.
Note: This figure needs to be updated!
The 4 corresponding differential equations are:
dR1/dt = k_syn_R1 * rep(S) * rep(R2) * rep(R3) - k_deg_R1 * R1
dR2/dt = k_syn_R2 * act(S) * rep(R3) * rep(R4) - k_deg_R2 * R2
dR3/dt = k_syn_R3 * rep(S) * rep(R1) * rep(R4) - k_deg_R3 * R3
dR4/dt = k_syn_R4 * act(S) * rep(R1) * rep(R2) - k_deg_R4 * R4
\_________________ _________________/ \_______ _______/
V V
synthesis rate degradation rate
where act(A) = (A/K_act)^n / (1+(A/K_act)^n)
and rep(R) = 1 / (1+(R/K_rep)^n)
K are affinity constants, while k are kinetic constants.
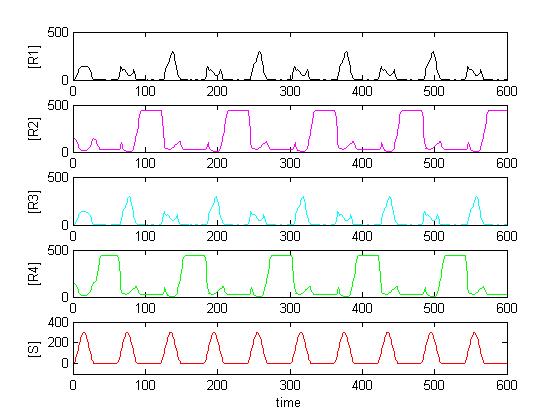
Solving the system in Matlab using reasonable affinity and kinetic constants, the result can look as in [Figure Simulation 2].
As expected, [R4] follows every second peak of [S].
By exploring the parameter space and performing Sensitivity Analysis, the following conclusions could be drawn:
- Changes in the affinity/cooperativity of the zink fingers affect the system more strongly than changes in the affinity/cooperativity of the input S.
- The degradation rate of the zinc fingers is the most sensitive parameter.
- The affinity constants R1..R4 should be as symmetrical as possible, in particular the couples R1,R3 and R2,R4. A difference in the affinity constant up to an order of magnitude appears tolerable.
- ...
More details of the simulation work are reported on the page Mathematical_Modeling.
Results
We have ongoing experiments and the documentation is not up to date / well structured. In the meantime, please refer to the Experiments page.
Discussion
Conclusion and Outlook
Appendix
References
atkinson03, basu05, bates05, Beerli00 (Linker), Beerli02, Beerli98, bulter04, cho04a-sensitivity, Chou et al. 1998, Dreier01 (ANN-paper), Dreier05 (CNN-paper), goryachev05, Greisman97, Isalan01, keiler01, Klug05 (Minireview), Lai04, Mani05, miller01, Newman03 (Leucine Zippers), Park et al. 2005 (Activation of transcription), römling02, ross91, Segal03, Segal99 (GNN-paper), sudesh00, suetsugu03, sutherland01, Yang95 (Kinetics!!), you04, zogaj01
Glossary
Previous Ideas
This is the brainstorming and previous ideas section. In this section you will find other projects that had been pursued, as well as random ideas without too much consideration of feasibility, etc.